Coronaviruses
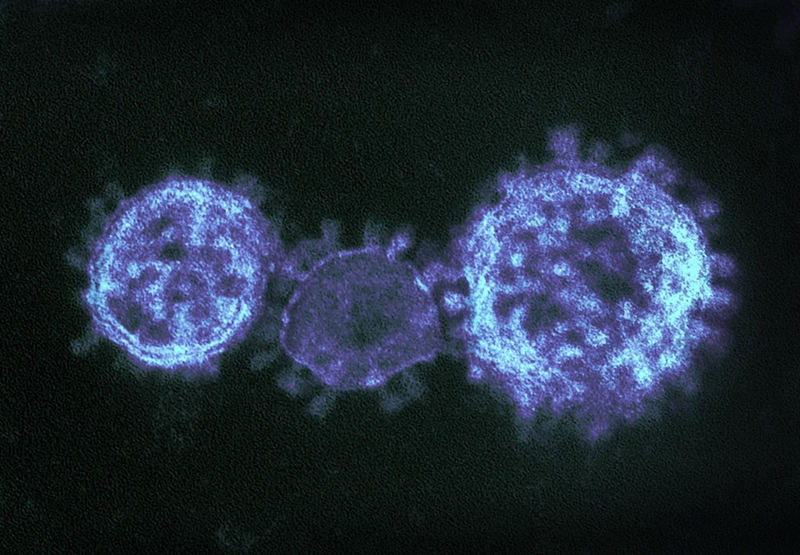
Coronaviruses are a large family of viruses that can infect a range of hosts. They are known to cause diseases including the common cold, Severe Acute Respiratory Syndrome (SARS) and Middle East Respiratory Syndrome (MERS) in humans.
In January 2020, China saw an outbreak of a new coronavirus strain now named SARS-CoV-2. Although the animal reservoir for the SARS and MERS viruses are known, this has yet to have been confirmed for SARS-CoV-2. All three strains are transmissible between humans.
To allow the widest possible distribution of relevant research, the Microbiology Society has brought together articles from across our portfolio and made this content freely available.
Image credit: "MERS-CoV" by NIAID is licensed under CC BY 2.0, this image has been modified.
Collection Contents
21 - 26 of 26 results
-
-
Primary structure of the S peplomer gene of bovine coronavirus and surface expression in insect cells
More LessThe nucleotide sequence of the S peplomer gene of bovine coronavirus (BCV) has been determined. A single open reading frame of 4089 nucleotides encodes a polypeptide of 150K with 20 potential sites for addition of N-linked oligosaccharides. Expression of the cloned BCV S gene by a recombinant of Autographa californica nuclear polyhedrosis virus resulted in production of a 180K glycosylated polypeptide which was transported to the surface of the cell. Comparison of the BCV S gene with the analogous genes of murine hepatitis viruses shows that the BCV S polypeptide contains a unique domain of 138 amino acids not present in murine hepatitis virus strain JHM, but which has a partially homologous counterpart in strain A59. This domain accounts for most of the differences in size of the S gene products of these coronaviruses.
-
-
-
The Peplomer Protein E2 of Coronavirus JHM as a Determinant of Neurovirulence: Definition of Critical Epitopes by Variant Analysis
More LessSummaryWe selected murine coronavirus JHM variants specifically changed in defined antigenic sites of the peplomer protein E2. Variants were isolated from the supernatants of monoclonal antibody hybridoma cell cultures which continued to secrete neutralizing antibodies after being infected with JHM. Comparative antigenic analysis and biological tests were performed in order to refine an operational epitope map and to characterize functional domains important for pathogenicity. The reaction patterns (neutralization, inhibition of cell fusion, immunofluorescence and binding in ELISA) between the variant viruses and the panel of monoclonal antibodies were very similar. Four groups of variants were characterized each of which revealed distinct changes affecting one defined antigenic site. These observations indicated that at least four independently mutable antigenic sites were associated with domains involved in cell fusion, neutralization and pathogenicity (E2-Aa, -Ab, -Ba and -Bb). JHM variants with alterations in the E2-Aa, -Ab or -Bb sites were similar to wild-type virus. These variants caused acute hepatitis and encephalomyelitis in mice. In contrast, JHM variants with changes in site E2-Ba had a strong propensity to induce chronic disease accompanied by demyelination persisting for several months.
-
-
-
The Predicted Primary Structure of the Peplomer Protein E2 of the Porcine Coronavirus Transmissible Gastroenteritis Virus
More LessSummaryThe complete nucleotide sequence of cloned cDNAs containing the E2 glycoproteinencoding region of the genome of transmissible gastroenteritis virus (TGEV) has been determined. A single large translatable frame of 4·3 kb starting at 8·2 kb from the 3′ end of the genome was identified. Its deduced amino acid sequence contains the characteristic features of a coronavirus peplomer protein: (i) the precursor polypeptide of TGEV E2 is 1447 residues long (i.e. 285 longer than the avian infectious bronchitis coronavirus spike protein); (ii) partial N-terminal sequencing demonstrated that a putative secretory signal sequence of 16 amino acids is absent in the virion-associated protein; (iii) the predicted mol. wt. of the apoprotein is 158K; most of the 32 potential N-glycosylation sites available in the sequence are presumed to be functional to account for the difference between this and the experimentally determined value (200K to 220K); (iv) a typical hydrophobic sequence near the C terminus is likely to be responsible for anchoring the peplomer to the virion envelope.
-
-
-
Plaque Assay for Titration of Bovine Enteric Coronavirus
More LessSUMMARYThe plaquing ability of two isolates of bovine enteric coronavirus (BECV) was studied in HRT18 (human rectal adenocarcinoma) cell monolayers. Both isolates were able to induce plaque formation within 2 to 3 days; plaques appeared as round opalescent areas which remained colourless after neutral red or crystal violet staining. A good correlation was found between the titres as determined either by counting the plaques that were visible to the naked eye before and after neutral red staining, or by enumerating fluorescence or haemadsorption foci.
-
-
-
The Polypeptide Structure of Canine Coronavirus and its Relationship to Porcine Transmissible Gastroenteritis Virus
More LessSUMMARYCanine coronavirus (CCV) isolate 1–71 was grown in secondary dog kidney cells and purified by rate zonal centrifugation. Polyacrylamide gel electrophoresis revealed four major structural polypeptides with apparent mol. wt. of 203800 (gp204), 49800 (p50), 31800 (gp32) and 21600 (gp22). Incorporation of 3H-glucosamine into gp204, gp32 and gp22 indicated that these were glycopolypeptides. Comparison of the structural polypeptides of CCV and porcine transmissible gastroenteritis virus (TGEV) by co-electrophoresis demonstrated that TGEV polypeptides corresponded closely, but not identically, with gp204, p50 and gp32 of CCV and confirmed that gp22 was a major structural component only in the canine virus. The close similarities in structure of the two coronaviruses augments the relationship established by serology.
-
-
-
Presence of Genomic Polyadenylate and Absence of Detectable Virion Transcriptase in Human Coronavirus OC-43
More LessSUMMARYHuman coronavirus RNA, prepared by extraction of purified virions with phenol-chloroform, consists of a major 15 to 55S class and a minor 4S class of RNA fragments. Polyadenylic acid [poly (A)] sequences are present in 15 to 55S but not in 4S RNA, suggesting different functions for each class. A stretch of poly (A) of approximately 19 adenosine monophosphate residues was obtained in sizing experiments after digesting OC-43 RNA with pancreatic and T1 ribonucleases. An OC-43 virion RNA transcriptase could not be detected with systems optimal for detecting the transcriptases of influenza and Newcastle disease virus.
-
