
Full text loading...
Attempts to identify members of the antiporter complement of the alkali- and saline-adapted soda lake alkaliphile Alkalimonas amylolytica N10 have used screens of DNA libraries in antiporter-deficient Escherichia coli KNabc. Earlier screens used Na+ or Li+ for selection but only identified one NhaD-type antiporter whose properties were inconsistent with a robust role in pH homeostasis. Here, new screens using elevated pH for selection identified three other putative antiporter genes that conferred resistance to pH ≥8.5 as well as Na+ resistance. The three predicted gene products were in the calcium/cation antiporter (CaCA), cation/proton antiporter-2 (CPA2) and cation/proton antiporter-1 (CPA1) families of membrane transporters, and were designated Aa-CaxA, Aa-KefB and Aa-NhaP respectively, reflecting homology within those families. Aa-CaxA conferred the poorest Na+ resistance and also conferred modest Ca2+ resistance. Aa-KefB and Aa-NhaP inhibited growth of a K+ uptake-deficient E. coli mutant (TK2420), suggesting that they catalysed K+ efflux. For Aa-NhaP, the reversibility of the growth inhibition by high K+ concentrations depended upon an organic nitrogen source, e.g. glutamine, rather than ammonium. This suggests that 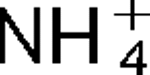 as well as K+ efflux is catalysed by Aa-NhaP. Vesicles of E. coli KNabc expressing Aa-NhaP, which conferred the strongest alkali resistance, exhibited K+/H+ antiport activity in a pH range from 7.5 to 9.5, and with an apparent K
m for K+ of 0.5 mM at pH 8.0. The properties of this antiporter are consistent with the possibility that this soda lake alkaliphile uses K+(
as well as K+ efflux is catalysed by Aa-NhaP. Vesicles of E. coli KNabc expressing Aa-NhaP, which conferred the strongest alkali resistance, exhibited K+/H+ antiport activity in a pH range from 7.5 to 9.5, and with an apparent K
m for K+ of 0.5 mM at pH 8.0. The properties of this antiporter are consistent with the possibility that this soda lake alkaliphile uses K+(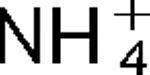 )/H+ antiport as part of its alkaline pH homeostasis mechanism and part of its capacity to reduce potentially toxic accumulation of cytoplasmic K+ or
)/H+ antiport as part of its alkaline pH homeostasis mechanism and part of its capacity to reduce potentially toxic accumulation of cytoplasmic K+ or 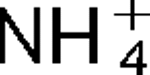 respectively, under conditions of high osmolarity or active amino acid catabolism.
respectively, under conditions of high osmolarity or active amino acid catabolism.

Article metrics loading...

Full text loading...
References


Data & Media loading...
