 -
Volume 60,
Issue 2,
2010
-
Volume 60,
Issue 2,
2010
Volume 60, Issue 2, 2010
- New Taxa
-
- Firmicutes And Related Organisms
-
-
Virgibacillus byunsanensis sp. nov., isolated from a marine solar saltern
More LessA Gram-variable, motile, endospore-forming and rod-shaped bacterial strain, ISL-24T, was isolated from a marine solar saltern of the Yellow Sea, Korea, and its taxonomic position was investigated by a polyphasic study. Strain ISL-24T grew optimally at pH 7.0–8.0, at 30–37 °C and in the presence of 8 % (w/v) NaCl. It contained MK-7 as the predominant menaquinone and anteiso-C15 : 0 as the predominant fatty acid. The DNA G+C content was 37.6 mol%. A phylogenetic analysis based on 16S rRNA gene sequences showed that strain ISL-24T fell within the genus Virgibacillus, clustering with Virgibacillus carmonensis LMG 20964T and Virgibacillus necropolis LMG 19488T, with a bootstrap resampling value of 92.3 %, and exhibiting 97.3 and 97.4 % 16S rRNA gene sequence similarity, respectively, to these strains. Strain ISL-24T exhibited 94.8–96.8 % 16S rRNA gene sequence similarity to the type strains of the other Virgibacillus species. Mean DNA–DNA relatedness values between strain ISL-24T and V. carmonensis DSM 14868T and V. necropolis DSM 14866T were 11 and 19 %, respectively. Differential phenotypic properties of strain ISL-24T, together with the phylogenetic and genetic distinctiveness, revealed that this strain is different from recognized Virgibacillus species. On the basis of phenotypic, phylogenetic and genetic data, strain ISL-24T represents a novel species of the genus Virgibacillus, for which the name Virgibacillus byunsanensis sp. nov. is proposed. The type strain is ISL-24T (=KCTC 13259T =CCUG 56754T).
-
-
-
Clostridium hydrogeniformans sp. nov. and Clostridium cavendishii sp. nov., hydrogen-producing bacteria from chlorinated solvent-contaminated groundwater
More LessFour hydrogen-producing, aerotolerant, anaerobic bacterial strains isolated from chlorinated solvent-contaminated groundwater were characterized using a polyphasic approach. Three of the strains, designated BL-18, BL-19 and BL-20T, were found to be identical in 16S rRNA gene sequences and in phenotypic properties. Cells of these strains are Gram-positive-staining, spore-forming, motile rods with peritrichous flagella. Growth occurred at 15–40 °C, pH 5.0–10.0 and at NaCl concentrations up to 5 % (w/v). Acid was produced in fermentation of cellobiose, fructose, galactose (weak), glucose, maltose and salicin. Products of fermentation in PYG medium were acetate, butyrate, ethanol, formate, carbon dioxide and hydrogen. Dominant cellular fatty acids when grown in PYG medium were C13 : 0 iso, C16 : 0, C13 : 0 anteiso, C15 : 0 iso and C15 : 0 anteiso. The genomic DNA G+C content was 30.4 mol%. These isolates can be differentiated from their closest phylogenetic relative, the cluster I Clostridium species Clostridium frigidicarnis (97.2 % similar to the type strain in 16S rRNA gene sequence), on the basis of phenotypic and chemotaxonomic properties. The other strain characterized in this study, BL-28T, was Gram-positive-staining with spore-forming, rod-shaped cells. Growth occurred at 15–46 °C, pH 6.0–8.5 and at NaCl concentrations up to 3 % (w/v). Acid was produced from cellobiose, dextran, fructose (weak), glucose, maltose, salicin and trehalose. End products of PYG fermentation included acetate, butyrate, pyruvate, carbon dioxide and hydrogen. Dominant cellular fatty acids from cells grown in PYG medium at 30 °C were C14 : 0, C14 : 0 dimethyl aldehyde, C16 : 0 and C12 : 0. The DNA G+C content was 28.5 mol%. Phylogenetic analysis based on 16S rRNA gene sequences revealed that strain BL-28T falls within cluster I of the genus Clostridium, but with ≤95.2 % identity with previously described species. On the basis of results presented here, strains BL-20T (=NRRL B-51348T =DSM 21757T) and BL-28T (=NRRL B-51352T =DSM 21758T) are proposed as the type strains of novel species of the genus Clostridium with the names Clostridium hydrogeniformans sp. nov. and Clostridium cavendishii sp. nov., respectively.
-
-
-
Fontibacillus aquaticus gen. nov., sp. nov., isolated from a warm spring
More LessA novel facultatively anaerobic strain, designated GPTSA 19T, was isolated from a warm spring and characterized using a polyphasic approach. The strain behaved as Gram-negative in the Gram staining procedure but showed a Gram-positive reaction in the aminopeptidase test. The novel strain was a mesophilic rod with ellipsoidal endospores. On the basis of 16S rRNA gene sequence analysis, the strain showed closest similarity (96.0 %) with Paenibacillus motobuensis MC10T. The gene sequence similarity of the novel strain with other species of the genus Paenibacillus was <95.8 %. The novel strain also had PAEN 515F and 682F signature sequence stretches in the 16S rRNA gene that are usually found in most species of the genus Paenibacillus. The strain possessed anteiso-C15 : 0 as the major fatty acid and MK-7 as the predominant menaquinone. Polar lipids included diphosphatidylglycerol (DPG), phosphatidylglycerol (PG), phosphatidylethanolamine (PE), six unknown phospholipids (PLs), one aminophospholipid (PN), three glycolipids (GLs), two aminolipids (ALs), one aminophosphoglycolipid (APGL) and three unknown lipids (ULs). The polar lipid profile of the novel strain, especially as regards ALs, GLs and PLs, distinguished it from the recognized type species of the genus Paenibacillus, Paenibacillus polymyxa, as well as from its closest relative P. motobuensis. Based on phenotypic and chemotaxonomic characteristics and analysis of the 16S rRNA gene sequence, the new strain merits the rank of a novel genus for which the name Fontibacillus gen. nov. is proposed. The type species of the new genus is Fontibacillus aquaticus gen. nov., sp. nov. with the type strain GPTSA 19T (=MTCC 7155T=DSM 17643T).
-
-
-
Alkalibacillus flavidus sp. nov., isolated from a marine solar saltern
More LessA Gram-stain-positive, motile, rod-shaped bacterial strain, ISL-17T, was isolated from a marine solar saltern of the Yellow Sea, Korea, and its taxonomic position was investigated by means of a polyphasic study. Strain ISL-17T grew optimally at pH 8.5–9.0, at 37 °C and in the presence of approximately 10 % (w/v) NaCl. It contained meso-diaminopimelic acid as the diagnostic diamino acid in the peptidoglycan, MK-7 as the predominant menaquinone and iso-C15 : 0, anteiso-C15 : 0 and iso-C16 : 0 as the major fatty acids. The DNA G+C content was 48.1 mol%. Phylogenetic analysis based on 16S rRNA gene sequences showed that strain ISL-17T fell within the genus Alkalibacillus, clustering with Alkalibacillus salilacus BH163T with a bootstrap resampling value of 100 %. Strain ISL-17T exhibited 98.2 % 16S rRNA gene sequence similarity to A. salilacus BH163T and 95.8–96.5 % similarity to the type strains of the other Alkalibacillus species. The mean DNA–DNA relatedness value between strain ISL-17T and A. salilacus KCTC 3916T was 19 %. The phenotypic properties of strain ISL-17T, together with its phylogenetic and genetic distinctiveness, enable this strain to be differentiated from recognized Alkalibacillus species. On the basis of phenotypic, phylogenetic and genetic data, strain ISL-17T represents a novel species within the genus Alkalibacillus, for which the name Alkalibacillus flavidus sp. nov. is proposed; the type strain is ISL-17T (=KCTC 13258T =CCUG 56753T).
-
- Other Bacteria
-
-
Bryobacter aggregatus gen. nov., sp. nov., a peat-inhabiting, aerobic chemo-organotroph from subdivision 3 of the Acidobacteria
More LessBryobacter aggregatus gen. nov., sp. nov. is proposed to accommodate three strains of slowly growing, chemo-organotrophic bacteria isolated from acidic Sphagnum peat bogs. These bacteria were strictly aerobic, Gram-negative, colourless, non-motile coccoids or short rods that multiplied by normal cell division and formed irregularly shaped cell aggregates. Strains MPL3T, MPL1011 and MOB76 were acidotolerant, mesophilic organisms capable of growth at pH 4.5–7.2 and between 4 and 33 °C (optimum growth at pH 5.5–6.5 and 22–28 °C). The preferred growth substrates were sugars, some heteropolysaccharides and galacturonic and glucuronic acids, which are released during decomposition of Sphagnum moss. The major fatty acids were iso-C15 : 0, C16 : 0 and summed feature 3 (iso-C15 : 0 2-OH and/or C16 : 1 ω7c); the major quinones were MK-9 and MK-10. The DNA G+C content was 55.5–56.5 mol%. Strains MPL3T, MPL1011 and MOB76 possessed nearly identical 16S rRNA gene sequences and belonged to the phylum Acidobacteria. They represent the first taxonomically characterized members of acidobacterial subdivision 3 and display only 81.7–86.7 % 16S rRNA gene sequence similarity to other members of the Acidobacteria with validly published names. Therefore, strains MPL3T, MPL1011 and MOB76 are classified as representatives of a novel species in a new genus, for which the name Bryobacter aggregatus gen. nov., sp. nov. is proposed; strain MPL3T (=ATCC BAA-1390T =DSM 18758T) is the type strain of Bryobacter aggregatus.
-
-
-
Thermocrinis minervae sp. nov., a hydrogen- and sulfur-oxidizing, thermophilic member of the Aquificales from a Costa Rican terrestrial hot spring
More LessA thermophilic bacterium, designated strain CR11T, was isolated from a filamentous sample collected from a terrestrial hot spring on the south-western foothills of the Rincón volcano in Costa Rica. The Gram-negative cells are approximately 2.4–3.9 μm long and 0.5–0.6 μm wide and are motile rods with polar flagella. Strain CR11T grows between 65 and 85 °C (optimum 75 °C, doubling time 4.5 h) and between pH 4.8 and 7.8 (optimum pH 5.9–6.5). The isolate grows chemolithotrophically with S0,
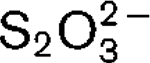 or H2 as the electron donor and with O2 (up to 16 %, v/v) as the sole electron acceptor. The isolate can grow on mannose, glucose, maltose, succinate, peptone, Casamino acids, starch, citrate and yeast extract in the presence of oxygen (4 %) and S0. Growth occurs only at NaCl concentrations below 0.4 % (w/v). The G+C content of strain CR11T is 40.3 mol%. Phylogenetic analysis of the 16S rRNA gene sequence places the strain as a close relative of Thermocrinis ruber OC 1/4T (95.7 % sequence similarity). Based on phylogenetic and physiological characteristics, we propose the name Thermocrinis minervae sp. nov., with CR11T (=DSM 19557T =ATCC BAA-1533T) as the type strain.
or H2 as the electron donor and with O2 (up to 16 %, v/v) as the sole electron acceptor. The isolate can grow on mannose, glucose, maltose, succinate, peptone, Casamino acids, starch, citrate and yeast extract in the presence of oxygen (4 %) and S0. Growth occurs only at NaCl concentrations below 0.4 % (w/v). The G+C content of strain CR11T is 40.3 mol%. Phylogenetic analysis of the 16S rRNA gene sequence places the strain as a close relative of Thermocrinis ruber OC 1/4T (95.7 % sequence similarity). Based on phylogenetic and physiological characteristics, we propose the name Thermocrinis minervae sp. nov., with CR11T (=DSM 19557T =ATCC BAA-1533T) as the type strain.
-
- Proteobacteria
-
-
Leisingera nanhaiensis sp. nov., isolated from marine sediment
More LessAn aerobic, Gram-staining-negative, motile, rod-shaped bacterium, strain NH52FT, was isolated from a sandy sediment sample taken from the South China Sea. On M2 agar medium (a complex medium), colonies were beige in colour. The isolate showed highest 16S rRNA gene sequence similarities to members of the genera Leisingera (96.7 % similarity), Phaeobacter (95.4–96.0 %) and Marinovum (94.1 %). Phylogenetic analysis based on 16S rRNA gene sequences indicated that strain NH52FT formed a distinct cluster with Leisingera methylohalidivorans MB2T and Leisingera aquimarina LMG 24366T. Optimal growth was observed at pH 7.0-8.5 and 25 °C and the new isolate required the presence of 1–4 % (w/v) NaCl. The major fatty acids were C18 : 1 ω7c, C16 : 0 2-OH, C10 : 0 3-OH, C12 : 0 3-OH, C16 : 0 and 11-methyl C18 : 1 ω7c. The DNA G+C content was 60.5 mol%. The phylogenetic and chemotaxonomic characteristics of strain NH52FT were similar to those of the genus Leisingera. However, the differences in phenotypic properties and the 16S rRNA gene similarity values demonstrated that the new isolate differed from recognized species of the genus Leisingera. On the basis of phenotypic, chemotaxonomic and phylogenetic data, this organism should be classified as a representative of a novel species in the genus Leisingera, for which the name Leisingera nanhaiensis sp. nov. is proposed. The type strain is NH52FT (=LMG 24841T=CCTCC AB 208316T=MCCC 1A04178T).
-
-
-
Altererythrobacter marensis sp. nov., isolated from seawater
More LessA novel Gram-negative bacterium, designated MSW-14T, was isolated from seawater collected around Mara Island, Jeju, Republic of Korea. The organism was motile by means of a flagellum and showed optimum growth at 0–4 % NaCl, 30 °C and pH 7.1. Phylogenetic analyses based on 16S rRNA gene sequences showed that the isolate belonged to the family Erythrobacteraceae. The strain's closest phylogenetic neighbours were Altererythrobacter epoxidivorans JCS350T (97.7 % sequence similarity), Altererythrobacter luteolus SW-109T (97.3 %) and Altererythrobacter indicus MSSRF26T (95.0 %). The dominant cellular fatty acid was C18 : 1 (summed feature 7, 52.8 %). The major ubiquinone was UQ-10. The DNA G+C content was 63.1 mol%. DNA–DNA hybridization values between strain MSW-14T and A. epoxidivorans KCCM 42314T and A. luteolus KCTC 12311T were 26.0–27.3 % and 9.8–15.2 %, respectively. On the basis of the data from the polyphasic characterization, strain MSW-14T represents a novel species, for which the name Altererythrobacter marensis sp. nov. is proposed. The type strain is MSW-14T (=KCTC 22370T=DSM 21428T).
-
-
-
Aestuariibacter litoralis sp. nov., isolated from a sandy sediment of the Sea of Japan
More LessThe phenotypic and phylogenetic characteristics of an aerobic, Gram-negative, motile, non-pigmented Alteromonas-like bacterium (designated strain KMM 3894T), isolated from a sandy sediment sample collected offshore of the Sea of Japan, were investigated. Comparative 16S rRNA gene sequence analysis revealed that strain KMM 3894T belonged to the genus Aestuariibacter and was most closely related to Aestuariibacter halophilus JC2043T (95.5 % sequence similarity). Fatty acid analysis showed C16 : 1 ω7c, C18 : 1 ω7c, and C16 : 0 as the dominant components. Strain KMM 3894T could be differentiated from recognized species of the genus Aestuariibacter by its ability to grow at 4 °C and at 30 °C, the optimum temperature for growth, and its inability to utilize most carbohydrates. On the basis of the phenotypic, chemotaxonomic and phylogenetic data, strain KMM 3894T is considered to represent a novel species of the genus Aestuariibacter, for which the name Aestuariibacter litoralis sp. nov. is proposed. The type strain is KMM 3894T (=NRIC 0754T=JCM 15896T).
-
-
-
Ochrobactrum pituitosum sp. nov., isolated from an industrial environment
More LessStrain CCUG 50899, a Gram-negative, rod-shaped, non-spore-forming, motile bacterium isolated from industrial environment in Sweden and tentatively assigned to the species Ochrobactrum anthropi, was studied in order to clarify its taxonomic status. 16S rRNA gene sequence similarities placed the strain in the genus Ochrobactrum, sharing highest similarity with the type strains of Ochrobactrum rhizosphaerae (99.3 %), Ochrobactrum thiophenivorans (98.7 %), Ochrobactrum pseudogrignonense (98.6 %) and Ochrobactrum grignonense (98.5 %). The fatty acid profile of [O. anthropi] CCUG 50899 (major fatty acids C18 : 1 ω7c and C19 : 0 cyclo ω8c and presence of C18 : 1 2-OH), the polar lipid profile (diphosphatidylglycerol, phosphatidylglycerol, phosphatidylmonomethylethanolamine, phosphatidylethanolamine, two unknown aminolipids and an unknown phospholipid), the presence of the quinone system ubiquinone Q-10 and a polyamine pattern with the major compounds putrescine and spermidine and moderate amounts of sym-homospermidine supported its affiliation to the genus Ochrobactrum. DNA–DNA reassociation experiments with the type strains of its closest relatives O. rhizosphaerae, O. pseudogrignonense, O. thiophenivorans and O. grignonense demonstrated that [O. anthropi] CCUG 50899 should be placed in a novel species, which is distinguishable from related species by a set of biochemical traits. Based on these data, reclassification of [O. anthropi] CCUG 50899 as the type strain of a novel species appears to be justified. Hence, we describe a novel species to accommodate this strain, for which we propose the name Ochrobactrum pituitosum sp. nov. The type strain is CCUG 50899T (=DSM 22207T).
-
-
-
Xenophilus aerolatus sp. nov., isolated from air
More LessA novel aerobic, Gram-negative, motile, rod-shaped bacterial strain designated 5516S-2T was isolated from an air sample taken in Suwon, Republic of Korea. Colonies were yellow-pigmented and circular with entire margins. 16S rRNA gene sequence analysis showed that strain 5516S-2T was closely related to Xylophilus ampelinus DSM 7250T (97.6 % sequence similarity), Variovorax soli KACC 11579T (97.5 %) and Xenophilus azovorans DSM 13620T (97.1 %). However, the phylogenetic tree indicated that strain 5516S-2T formed a separate clade from Xenophilus azovorans. Strain 5516S-2T displayed 42, 31 and 30 % DNA–DNA relatedness to the type strains of Xenophilus azovorans, Xylophilus ampelinus and V. soli, respectively. The major fatty acids (>10 % of total fatty acids) were C16 : 0 (33.3 %), C17 : 0 cyclo (18.8 %), C18 : 1 ω7c (17.5 %) and summed feature 3 (comprising C16 : 1 ω7c and/or iso-C15 : 0 2-OH; 13.0 %). The DNA G+C content was 69 mol%. The major quinone was ubiquinone Q-8. The predominant polar lipids were diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylglycerol and two unknown aminophospholipids. Genotypic and phenotypic characteristics clearly distinguished strain 5516S-2T from closely related species and indicated that it represents a novel species within the genus Xenophilus, for which the name Xenophilus aerolatus sp. nov. is proposed. The type strain is 5516S-2T (=KACC 12602T=DSM 19424T).
-
-
-
Halomonas xinjiangensis sp. nov., a halotolerant bacterium isolated from a salt lake
More LessA novel bacterium, TRM 0175T, belonging to the genus Halomonas, was isolated from a soil sample taken from a salt lake in Xinjiang Province, north-west China. The isolate was Gram-negative, aerobic, rod-shaped and motile by means of peritrichous flagella. It was catalase-positive and oxidase-negative. Growth occurred at NaCl concentrations of 0–20 % (optimum at 10–13 %), at 15–50 °C (optimum at 37 °C) and at pH 6.0–9.0 (optimum at pH 7.0). Metabolism was respiratory with oxygen as terminal electron acceptor. Acid was produced from d-ribose, d- and l-arabinose, d-xylose, d-galactose, d-mannose, l-rhamnose, cellobiose, maltose, trehalose and d- and l-fucose and was produced weakly from aesculin. The predominant ubiquinone was Q-9. The major fatty acids were C18 : 1 ω7c and C19 : 0 cyclo ω8c. The G+C content of the genomic DNA was 60.0 mol%. The affiliation of strain TRM 0175T with the genus Halomonas was confirmed by 16S rRNA gene sequence comparisons. The most closely related species was Halomonas anticariensis; 16S rRNA gene sequence similarity between H. anticariensis FP35T and strain TRM 0175T was 95.3 %. Phenotypically, some characteristics of TRM 0175T differed from those of H. anticariensis. On the basis of data from this polyphasic study, strain TRM 0175T represents a novel species of the genus Halomonas, for which the name Halomonas xinjiangensis sp. nov. is proposed; the type strain is TRM 0175T (=CCTCC AB 208329T =KCTC 22608T).
-
-
-
Halomonas stevensii sp. nov., Halomonas hamiltonii sp. nov. and Halomonas johnsoniae sp. nov., isolated from a renal care centre
More LessA total of 14 Halomonas strains were isolated from the blood of two patients and from dialysis machines of a renal care centre. The strains were Gram-negative, halophilic, motile and non-spore-forming rods. They produced cream-coloured colonies and contained Q-9 as the predominant ubiquinone and C18 : 1 ω7c and C16 : 0 as the major fatty acids. Phylogenetic analysis based on 16S rRNA gene sequencing showed that the 14 isolates were most closely related to Halomonas magadiensis 21 MIT with 98.1–98.9 % sequence similarity and that they formed three separate lineages among themselves. Combined phenotypic and DNA–DNA hybridization data support the conclusion that they represent three novel species of the genus Halomonas, for which the names Halomonas stevensii sp. nov. (type strain S18214T=KCTC 22148T=DSM 21198T), Halomonas hamiltonii sp. nov. (type strain W1025T=KCTC 22154T=DSM 21196T) and Halomonas johnsoniae sp. nov. (type strain T68687T=KCTC 22157T=DSM 21197T) are proposed.
-
-
-
Sphingobium faniae sp. nov., a pyrethroid-degrading bacterium isolated from activated sludge treating wastewater from pyrethroid manufacture
More LessA bacterial strain capable of degrading pyrethroid, designated JZ-2T, was isolated from activated sludge treating pyrethroid-contaminated wastewater. Phylogenetic analysis based on 16S rRNA gene sequences indicated that strain JZ-2T belongs to the genus Sphingobium. It showed the highest levels of 16S rRNA gene sequence similarity to Sphingobium cloacae JCM 10874T (98.3 %) and Sphingobium ummariense CCM 7431T (97.1 %), and 94.8–96.9 % similarity to the type strains of other members of the genus Sphingobium. Strain JZ-2T contained C18 : 1 ω7c as the predominant fatty acid, C14 : 0 2-OH as the major 2-hydroxy fatty acid, ubiquinone Q-10 as the main respiratory quinone, diphosphatidylglycerol, phosphatidylglycerol, phosphatidylcholine, phosphatidylmonomethylethanolamine, phosphatidylethanolamine and two sphingoglycolipids as the predominant polar lipids and spermidine as the major polyamine. DNA–DNA hybridization results showed that strain JZ-2T had low genomic relatedness with S. cloacae JCM 10874T (34 %) and S. ummariense CCM 7431T (23 %). Based on the phenotypic, genotypic and phylogenetic data presented, strain JZ-2T is considered to represent a novel species of the genus Sphingobium, for which the name Sphingobium faniae sp. nov. is proposed. The type strain is JZ-2T (=CGMCC 1.7749T =DSM 21829T).
-
-
-
Sphingobium scionense sp. nov., an aromatic hydrocarbon-degrading bacterium isolated from contaminated sawmill soil
More LessThis study characterized strain WP01T, a Gram-staining-negative, rod-shaped, aerobic bacterium isolated from a polycyclic aromatic hydrocarbon-contaminated soil in New Zealand. Strain WP01T shared many characteristics of the genus Sphingobium: the predominant respiratory quinone (89 %) was ubiquinone with ten isoprene units (Q-10); the major fatty acids were C18 : 1 ω7c, C16 : 1 ω7c, C16 : 0 and C14 : 0 2-OH; spermidine was the major polyamine; the DNA G+C content was 63.8 mol%; and the Sphingobium-specific 16S rRNA signatures were conserved. A point of difference from other species of the genus Sphingobium was that strain WP01T reduced nitrate to nitrite. The polar lipid pattern consisted of the predominant compounds diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylglycerol and sphingoglycolipids. 16S rRNA gene sequence analysis showed that, amongst the recognized species of the genus Sphingobium, strain WP01T was most similar to Sphingobium yanoikuyae GIFU 9882T and Sphingobium amiense YTT (>97 % 16S rRNA gene sequence similarities). The low DNA–DNA relatedness values between strain WP01T and S. yanoikuyae GIFU 9882T (46.6 %) and S. amiense DSM 16289T (25.6 %) indicated no relatedness at the species level. On the basis of these characteristics, it is concluded that strain WP01T should be considered as representing a novel species within the genus Sphingobium, for which the name Sphingobium scionense sp. nov. is proposed. The type strain is WP01T (=DSM 19371T=ICMP 13533T).
-
-
-
Sphingobium quisquiliarum sp. nov., a hexachlorocyclohexane (HCH)-degrading bacterium isolated from an HCH-contaminated soil
More LessA yellow-pigmented, hexachlorocyclohexane (HCH)-degrading bacterial strain, P25T, was isolated from an HCH dump site located in the northern part of India. Phylogenetic analysis based on the 16S rRNA gene sequence showed that the strain belongs to the genus Sphingobium, as it showed highest sequence similarity to Sphingobium amiense IAM 15006T (97.7 %). The 16S rRNA gene sequence similarity between strain P25T and members of other species of the genus Sphingobium with validly published names ranged from 94.0 to 97.7 %. The DNA–DNA relatedness between strain P25T and Sphingobium amiense IAM 15006T and other related strains was found be less than 30 %, confirming it to represent a novel species. The DNA G+C content of strain P25T was 65 mol%. The polyamine profile showed the presence of spermidine. The predominant cellular fatty acids were summed feature 8 (18 : 1ω7c and/or 18 : 1ω6c; 48.3 %), 16 : 0 (13.7 %) and 14 : 0 2-OH (8.8 %). The polar lipid profile of strain P25T also corresponded to those reported for sphingomonads (phosphatidylethanolamine, diphosphatidylglycerol, phosphatidyldimethylethanolamine, phosphatidylglycerol, phosphatidylcholine, sphingoglycolipid), supporting its identification as a member of the family Sphingomonadaceae. The results obtained from DNA–DNA hybridization and biochemical and physiological tests clearly distinguished strain P25T from closely related members of the genus Sphingobium. Thus, a novel species of the genus Sphingobium is proposed, Sphingobium quisquiliarum sp. nov. The type strain is P25T (=MTCC 9472T =CCM 7543T).
-
-
-
Thiohalobacter thiocyanaticus gen. nov., sp. nov., a moderately halophilic, sulfur-oxidizing gammaproteobacterium from hypersaline lakes, that utilizes thiocyanate
More LessA moderately halophilic, obligately chemolithoautotrophic, sulfur-oxidizing bacterium, designated strain HRh1T, was obtained from mixed sediment samples from hypersaline chloride–sulfate lakes in the Kulunda Steppe, in south-western Siberia (Russia), using aerobic enrichment culture at 1 M NaCl with thiocyanate as substrate. Cells of the isolate were short, non-motile rods with a Gram-negative type of cell wall. The bacterium was an obligate aerobe capable of chemolithoautotrophic growth with thiocyanate and thiosulfate. With thiosulfate, it grew at NaCl concentrations of 0.2–3.0 M (optimum 0.5 M) and at pH 6.3–8.0 (optimum pH 7.3–7.5). During growth on thiocyanate, cyanate was identified as an intermediate. The dominant cellular fatty acids were C16 : 0 and C18 : 1 ω7. Phylogenetic analysis based on 16S rRNA gene sequencing placed the isolate in the class Gammaproteobacteria as an independent lineage, with an unclassified marine sulfur-oxidizing bacterium as the closest culturable relative (93 % sequence similarity). A single cbbL gene (coding for the key enzyme of the Calvin–Benson cycle of autotrophic CO2 assimilation) with relatively low similarity to any homologous genes found in chemolithoautotrophs was detected in strain HRh1T. On the basis of our phenotypic and phylogenetic analysis, the halophilic isolate is proposed to represent a new genus and novel species, Thiohalobacter thiocyanaticus gen. nov., sp. nov. The type strain of Thiohalobacter thiocyanaticus is HRh1T (=DSM 21152T =UNIQEM U249T).
-
- Eukaryotic Micro-Organisms
-
-
Description of Eurystomatella sinica n. gen., n. sp., with establishment of a new family Eurystomatellidae n. fam. (Protista, Ciliophora, Scuticociliatia) and analyses of its phylogeny inferred from sequences of the small-subunit rRNA gene
More LessRecently, an undescribed marine ciliate was isolated from China. Investigation of its morphology and infraciliature revealed it as an undescribed species representing a new genus, Eurystomatella n. gen., the type of the new family Eurystomatellidae n. fam. The new family is defined by close-set, apically positioned oral membranelles and a dominant buccal field that is surrounded by an almost completely circular paroral membrane. The new genus is defined by having a small oral membranelle 1 (M1), bipartite M2 and well-developed M3, a body surface faintly sculptured with a silverline system in a quadrangular, reticulate pattern and a cytostome located at the anterior third of a large buccal field. The type species of the new genus, Eurystomatella sinica n. sp., is a morphologically unique form that is defined mainly by the combination of a conspicuously flattened body, several caudal cilia, extremely long cilia associated with the buccal apparatus and a contractile vacuole located subcaudally. According to phylogenetic analyses of small-subunit (SSU) rRNA gene sequences, Eurystomatella clusters with the genus Cyclidium, as a sister group to the family Pleuronematidae. The great divergence in both buccal and somatic ciliature between Eurystomatella and all other known scuticociliates supports the establishment of a new family for Eurystomatella.
-
Volumes and issues
-
Volume 74 (2024)
-
Volume 73 (2023)
-
Volume 72 (2022 - 2023)
-
Volume 71 (2020 - 2021)
-
Volume 70 (2020)
-
Volume 69 (2019)
-
Volume 68 (2018)
-
Volume 67 (2017)
-
Volume 66 (2016)
-
Volume 65 (2015)
-
Volume 64 (2014)
-
Volume 63 (2013)
-
Volume 62 (2012)
-
Volume 61 (2011)
-
Volume 60 (2010)
-
Volume 59 (2009)
-
Volume 58 (2008)
-
Volume 57 (2007)
-
Volume 56 (2006)
-
Volume 55 (2005)
-
Volume 54 (2004)
-
Volume 53 (2003)
-
Volume 52 (2002)
-
Volume 51 (2001)
-
Volume 50 (2000)
-
Volume 49 (1999)
-
Volume 48 (1998)
-
Volume 47 (1997)
-
Volume 46 (1996)
-
Volume 45 (1995)
-
Volume 44 (1994)
-
Volume 43 (1993)
-
Volume 42 (1992)
-
Volume 41 (1991)
-
Volume 40 (1990)
-
Volume 39 (1989)
-
Volume 38 (1988)
-
Volume 37 (1987)
-
Volume 36 (1986)
-
Volume 35 (1985)
-
Volume 34 (1984)
-
Volume 33 (1983)
-
Volume 32 (1982)
-
Volume 31 (1981)
-
Volume 30 (1980)
-
Volume 29 (1979)
-
Volume 28 (1978)
-
Volume 27 (1977)
-
Volume 26 (1976)
-
Volume 25 (1975)
-
Volume 24 (1974)
-
Volume 23 (1973)
-
Volume 22 (1972)
-
Volume 21 (1971)
-
Volume 20 (1970)
-
Volume 19 (1969)
-
Volume 18 (1968)
-
Volume 17 (1967)
-
Volume 16 (1966)
-
Volume 15 (1965)
-
Volume 14 (1964)
-
Volume 13 (1963)
-
Volume 12 (1962)
-
Volume 11 (1961)
-
Volume 10 (1960)
-
Volume 9 (1959)
-
Volume 8 (1958)
-
Volume 7 (1957)
-
Volume 6 (1956)
-
Volume 5 (1955)
-
Volume 4 (1954)
-
Volume 3 (1953)
-
Volume 2 (1952)
-
Volume 1 (1951)
Most Read This Month


