 -
Volume 60,
Issue 1,
2010
-
Volume 60,
Issue 1,
2010
Volume 60, Issue 1, 2010
- Editorial
-
- Validation List
-
-
-
List of new names and new combinations previously effectively, but not validly, published
More LessThe purpose of this announcement is to effect the valid publication of the following effectively published new names and new combinations under the procedure described in the Bacteriological Code (1990 Revision). Authors and other individuals wishing to have new names and/or combinations included in future lists should send three copies of the pertinent reprint or photocopies thereof, or an electronic copy of the published paper, to the IJSEM Editorial Office for confirmation that all of the other requirements for valid publication have been met. It is also a requirement of IJSEM and the ICSP that authors of new species, new subspecies and new combinations provide evidence that types are deposited in two recognized culture collections in two different countries. It should be noted that the date of valid publication of these new names and combinations is the date of publication of this list, not the date of the original publication of the names and combinations. The authors of the new names and combinations are as given below, and these authors' names will be included in the author index of the present issue. Inclusion of a name on these lists validates the publication of the name and thereby makes it available in bacteriological nomenclature. The inclusion of a name on this list is not to be construed as taxonomic acceptance of the taxon to which the name is applied. Indeed, some of these names may, in time, be shown to be synonyms, or the organisms may be transferred to another genus, thus necessitating the creation of a new combination.
-
-
- Notification List
-
-
-
Notification that new names and new combinations have appeared in volume 59, part 10, of the IJSEM
More LessThis listing of names published in a previous issue of the IJSEM is provided as a service to bacteriology to assist in the recognition of new names and new combinations. This procedure was proposed by the Judicial Commission [Minute 11(ii), Int J Syst Bacteriol 41 (1991), p. 185]. The names given herein are listed according to the Rules of priority (i.e. page number and order of valid publication of names in the original articles). Taxonomic opinions included in this List (i.e. the creation of synonyms or the emendation of circumscriptions) cannot be considered as validly published nor, in any other way, approved by the International Committee on Systematics of Prokaryotes and its Judicial Commission.
-
-
- List Of Changes In Taxonomic Opinion
-
-
-
Notification of changes in taxonomic opinion previously published outside the IJSEM
More LessThe Bacteriological Code deals with the nomenclature of prokaryotes. This may include existing names (the Approved Lists of Bacterial Names) as well as new names and new combinations. In this sense the Code is also dealing indirectly with taxonomic opinions. However, as with most codes of nomenclature there are no mechanisms for formally recording taxonomic opinions that do not involve the creation of new names or new combinations. In particular, it would be desirable for taxonomic opinions resulting from the creation of synonyms or emended descriptions to be made widely available to the public. In 2004, the Editorial Board of the International Journal of Systematic and Evolutionary Microbiology (IJSEM) agreed unanimously that it was desirable to cover such changes in taxonomic opinions (i.e. the creation of synonyms or the emendation of circumscriptions) previously published outside the IJSEM, and to introduce a List of Changes in Taxonomic Opinion [Notification of changes in taxonomic opinion previously published outside the IJSEM; Euzéby et al. (2004). Int J Syst Evol Microbiol 54, 1429–1430]. Scientists wishing to have changes in taxonomic opinion included in future lists should send one copy of the pertinent reprint or a photocopy or a PDF file thereof to the IJSEM Editorial Office or to the Lists Editor. It must be stressed that the date of proposed taxonomic changes is the date of the original publication not the date of publication of the list. Taxonomic opinions included in the List of Changes in Taxonomic Opinion cannot be considered as validly published nor, in any other way, approved by the International Committee on Systematics of Prokaryotes and its Judicial Commission. The names that are to be used are those that are the ‘correct names’ (in the sense of Principle 6) in the opinion of the bacteriologist, with a given circumscription, position and rank. A particular name, circumscription, position and rank does not have to be adopted in all circumstances. Consequently, the List of Changes in Taxonomic Opinion must be considered as a service to bacteriology and it has no ‘official character’, other than providing a centralized point for registering/indexing such changes in a way that makes them easily accessible to the scientific community.
-
-
- New Taxa
-
- Actinobacteria
-
-
Streptomyces artemisiae sp. nov., isolated from surface-sterilized tissue of Artemisia annua L.
More LessThe taxonomic position of an actinomycete strain YIM 63135T, which was isolated from the surface-sterilized tissue of Artemisia annua L. collected from Yunnan province, south-west China, was determined by using a polyphasic approach. Morphological and chemical characteristics of the novel strain were consistent with those of the genus Streptomyces. It developed a pinkish aerial mycelium and pinkish-brown substrate mycelium on oatmeal agar. The cell wall of the strain contained ll-diaminopimelic acid. The menaquinones comprised MK-9(H6) (62.8 %), MK-9(H8) (31.4 %) and MK-9(H4) (5.9 %). The phospholipids were diphosphatidylglycerol, phosphatidylglycerol, phosphatidylinositol, phosphatidylethanolamine, an unknown glucosamine-containing phospholipid (GluNu), phosphatidylinositol mannosides and four unknown ninhydrin-negative phospholipids. The major fatty acids were iso-C16 : 0 (30.0 %), anteiso-C17 : 0 (27.3 %) and anteiso-C15 : 0 (17.0 %). The DNA G+C content of strain YIM 63135T was 72.6 mol%. Phylogenetic analysis of 16S rRNA gene sequences revealed that strain YIM 63135T is a member of the genus Streptomyces and exhibited 99.9 % gene sequence similarity to Streptomyces armeniacus NBRC 12555T, while low sequence similarity values (<97.0 %) distinguished strain YIM 63135T from all other Streptomyces species. DNA–DNA hybridization studies suggested that strain YIM 63135T represents a different genomic species. On the basis of phenotypic and phylogenetic characteristics, strain YIM 63135T was considered to represent a novel species of the genus Streptomyces, for which the name Streptomyces artemisiae sp. nov. is proposed, with YIM 63135T (=CCTCC AA 208059T =DSM 41953T) as the type strain.
-
-
-
Alloactinosynnema album gen. nov., sp. nov., a member of the family Actinosynnemataceae isolated from soil
More LessThe taxonomic position of a Gram-stain-positive, aerobic strain, designated 03-9939T, isolated from a soil sample collected from Xinjiang Province, China, was established using a polyphasic approach. Whole-cell hydrolysates of strain 03-9939T contained galactose and ribose as diagnostic sugars and meso-diaminopimelic acid as the diamino acid. The predominant menaquinone was MK-9(H4). The phospholipids consisted of diphosphatidylglycerol, phosphatidylglycerol and phosphatidylcholine. The major fatty acids were iso-C16 : 0 (61.5 %) and iso-C16 : 1 H (11.6 %). The genomic DNA G+C content was 68.2 mol%. Phylogenetic analysis based on 16S rRNA gene sequences revealed that strain 03-9939T should be placed within the family Actinosynnemataceae, in which the strain formed a distinct lineage. Signature nucleotides in the 16S rRNA gene sequence showed that the strain contained a genus-specific diagnostic nucleotide signature pattern. The combination of phylogenetic analysis, phenotypic characteristics and chemotaxonomic data supported the conclusion that strain 03-9939T represents a novel species in a new genus of the family Actinosynnemataceae, for which the name Alloactinosynnema album gen. nov., sp. nov. is proposed. Strain 03-9939T (=DSM 45114T =KCTC 19294T =CCM 7461T) is the type strain of Alloactinosynnema album.
-
-
-
Actinopolymorpha cephalotaxi sp. nov., a novel actinomycete isolated from rhizosphere soil of the plant Cephalotaxus fortunei
More LessAn actinomycete, strain I06-2230T, was isolated from rhizosphere soil of the plant Cephalotaxus fortunei, collected from Yunnan province, south China. Phylogenetic analysis based on 16S rRNA gene sequences indicated that the isolate belongs to the genus Actinopolymorpha. Cells grew on agar surfaces, with no penetration even after prolonged cultivation. Aerial hyphae were absent. Cells were irregularly shaped and remained attached as chains or aggregates. Chemotaxonomic data, which showed ll-diaminopimelic acid in the cell wall, glucose as the whole-cell sugar, type PI phospholipids and MK-9(H4) as the predominant menaquinone, supported the affiliation of strain I06-2230T to the genus Actinopolymorpha. The major fatty acids were iso-C15 : 0, iso-C16 : 0 and iso-C16 : 1 H. The genomic DNA G+C content was 69.3 mol%. DNA–DNA hybridization data, in combination with chemotaxonomic, physiological and biochemical data, demonstrated that strain I06-2230T should be classified as representing a novel species of the genus Actinopolymorpha. The name Actinopolymorpha cephalotaxi sp. nov. is proposed, with strain I06-2230T (=DSM 45117T=CCM 7466T=KCTC 19293T) as the type strain.
-
-
-
Friedmanniella luteola sp. nov., Friedmanniella lucida sp. nov., Friedmanniella okinawensis sp. nov. and Friedmaniella sagamiharensis sp. nov., isolated from spiders
More LessFour Gram-positive, non-motile, aerobic actinobacteria were isolated from spiders and their webs. Their genetic, phenotypic and chemical properties were studied. The 16S rRNA gene sequence data suggested that the four novel isolates belonged to the genus Friedmanniella. Two strains (FA1T and FA2T) formed a cluster together with Friedmanniella capsulata and Friedmanniella lacustris and the other two strains (FB1T and FB2T) formed a cluster together with Friedmanniella antarctica and Friedmanniella spumicola. The cell-wall peptidoglycan contained ll-A2pm and mycolic acids were absent. Isoprenoid quinones were mainly composed of MK-9(H4), MK-9(H2) and MK-9 and the predominant fatty acids were 12-methyltetradecanoic acid (ai-C15 : 0) and 13-methyltetradecanoic acid (i-C15 : 0). The major polar lipids were phosphatidylinositol and phosphatidylglycerol. In addition, strain FA1T, FB1T, and FB2T contained diphosphatidylglycerol and phosphatidylcholine. The DNA G+C contents were: 72 mol%, 73 mol%, 74 mol% and 75 mol% for strains FA1T, FA2T, FB1T, and FB2T, respectively. DNA–DNA hybridization studies demonstrated that the novel strains showed low relatedness values to F. capsulata, F. lacustris, F. antarctica and F. spumicola. These data support the proposal that strains FA1T, FA2T, FB1T and FB2T represent novel species of the genus Friedmanniella. Therefore, the names Friedmanniella luteola (type strain FA1T=DSM 21741T=NBRC 104963T), Friedmanniella lucida (type strain FA2T=DSM 21742T=NBRC 104964T), Friedmanniella okinawensis (type strain FB1T=DSM 21744T=NBRC 104966T) and Friedmanniella sagamiharensis (type strain FB2T=DSM 21743T=NBRC 104965T) are proposed for these new strains.
-
-
-
Kocuria koreensis sp. nov., isolated from fermented seafood
More LessA Gram-positive, aerobic, non-motile and coccoid actinobacterium, designated P31T, was isolated from a traditional, fermented seafood. The strain was catalase-positive and oxidase-negative. Cells grew in the presence of 0–15.0 % (w/v) NaCl, and at pH 5–10 and 15–37 °C. Major cellular fatty acids were anteiso-C15 : 0, anteiso-C17 : 0 and iso-C16 : 0. Strain P31T contained MK-7 as the predominant menaquinone. The DNA G+C content of the genomic DNA of strain P31T was 65.2 mol%. A phylogenetic analysis based on the 16S rRNA gene sequence indicated that strain P31T was most closely related to Kocuria kristinae DSM 20032T, with 96.9 % similarity, and these two strains clustered together in constructed phylogenetic trees. The DNA–DNA hybridization value between strain P31T and K. kristinae DSM 20032T was 21.1 %. On the basis of the phenotypic, chemotaxonomic and phylogenetic data, it is suggested that strain P31T represents a novel species of the genus Kocuria, for which the name Kocuria koreensis sp. nov. is proposed. The type strain is P31T (=KCTC 19595T=JCM 15915T).
-
-
-
Actinomadura sputi sp. nov., isolated from the sputum of a patient with pulmonary infection
More LessThe taxonomic position of an actinomycete, strain IMMIB L-889T, isolated from the sputum of a 64-year-old man, was determined using a polyphasic taxonomic approach. The strain had chemical and morphological properties that were consistent with its classification in the genus Actinomadura. It formed a distinct phyletic line in the 16S rRNA gene tree of Actinomadura and was most closely related to the type strain of Actinomadura hallensis (98.4 % sequence similarity), but could be readily distinguished from the latter species using DNA–DNA relatedness and phenotypic data. The combined genotypic and phenotypic data indicate that strain IMMIB L-889T represents a novel species of the genus Actinomadura, for which the name Actinomadura sputi sp. nov. is proposed. The type strain is IMMIB L-889T (=DSM 45233T=CCUG 56587T).
-
- Bacteroidetes
-
-
Arenibacter nanhaiticus sp. nov., isolated from marine sediment of the South China Sea
More LessAn aerobic, Gram-stain-negative, rod-shaped bacterial isolate, strain NH36AT, was isolated from a sandy sediment sample from the South China Sea. Colonies of the isolate were dark orange on M2 agar. Optimal growth was observed at pH 7.0–8.5, 30 °C and in the presence of 0.5–4.0 % (w/v) NaCl. The major fatty acids were C15 : 0, iso-C15 : 0, anteiso-C15 : 0, iso-C15 : 1, iso-C15 : 0 3-OH, iso-C17 : 0 3-OH and summed feature 3 (comprising iso-C15 : 0 2-OH and/or C16 : 1 ω7c). The DNA G+C content was 38.9 mol%. 16S rRNA gene sequence analysis revealed that strain NH36AT was most closely related to members of the genus Arenibacter, exhibiting 94.3–96.2 % sequence similarity to the type strains of Arenibacter species. On the basis of phenotypic, chemotaxonomic and phylogenetic data, this organism should be classified as a representative of a novel species in the genus Arenibacter. The name Arenibacter nanhaiticus sp. nov. is proposed and the type strain is NH36AT (=LMG 24842T=CCTCC AB 208315T=MCCC 1A04137T).
-
-
-
Pontibacter xinjiangensis sp. nov., in the phylum ‘Bacteroidetes’, and reclassification of [Effluviibacter] roseus as Pontibacter roseus comb. nov.
More LessA novel strain, designated 311-10T, isolated from soil of Xinjiang, China, was characterized by using a polyphasic taxonomic approach. The isolate was Gram-negative, aerobic, rod-shaped, non-motile, oxidase-negative and catalase-positive. The predominant menaquinone of strain 311-10T was MK-7 and the genomic DNA G+C content was 47.8 mol%. Phylogenetic analysis based on 16S rRNA gene sequences indicated that the isolate formed a cluster with the genera Pontibacter and [Effluviibacter] in the phylum ‘Bacteroidetes’, with sequence similarities of 93.9–95.6 %. Phylogenetic evidence and the results of phenotypic, genotypic and chemotaxonomic analyses support the reclassification of [Effluviibacter] roseus as Pontibacter roseus comb. nov. (type strain, SRC-1T=MTCC 7260T=DSM 17521T) and the establishment of a novel species, Pontibacter xinjiangensis sp. nov., with strain 311-10T (=CCTCC AB 207200T=NRRL B-51335T) as the type strain.
-
-
-
Meridianimaribacter flavus gen. nov., sp. nov., a member of the family Flavobacteriaceae isolated from marine sediment of the South China Sea
More LessA Gram-staining-negative, rod-shaped marine bacterium, designated strain NH57NT, isolated from sandy sediment in the Mischief Reef of the South China Sea, was characterized based on its physiological and biochemical features, fatty acid profile and phylogenetic position. 16S rRNA gene sequence analysis revealed a clear affiliation with the family Flavobacteriaceae. Strain NH57NT showed the closest phylogenetic relationship with members of the genera Gaetbulibacter, Gelidibacter, Subsaxibacter, Subsaximicrobium and Yeosuana; levels of 16S rRNA gene sequence similarity between strain NH57NT and the type strains of related species ranged from 94.9 to 91.2 %. Cells of strain NH57NT were motile by gliding and grew on solid media as yellow colonies at 9–37 °C, pH 6.5–8.5 and in the presence of 0.5–4.0 % NaCl. The DNA G+C content was 32.7 mol% and the predominant fatty acids were iso-C15 : 1 (22.7 % of the total), iso-C15 : 0 (20.7 %), iso-C17 : 0 3-OH (9.5 %), iso-C16 : 0 3-OH (8.3 %), C15 : 0 (7.8 %) and iso-C15 : 0 3-OH (5.8 %). Based on the physiological and phylogenetic data, and on the fatty acid composition, strain NH57NT is considered to represent a novel species of a new genus in the family Flavobacteriaceae, for which the name Meridianimaribacter flavus gen. nov., sp. nov. is proposed. The type strain of Meridianimaribacter flavus is NH57NT (=CCTCC AB 208318T=LMG 24839T=MCCC 1A03544T).
-
-
-
Mucilaginibacter rigui sp. nov., isolated from wetland freshwater, and emended description of the genus Mucilaginibacter
More LessA non-motile, rod-shaped bacterium, designated strain WPCB133T, was isolated from freshwater collected from the Woopo wetland (Republic of Korea). Cells were Gram-reaction-negative, aerobic and catalase- and oxidase-positive. The major fatty acids were iso-C15 : 0 and anteiso-C15 : 0. The strain contained MK-7 as the major isoprenoid quinone. The DNA G+C content was 47 mol%. A phylogenetic tree based on 16S rRNA gene sequences showed that strain WPCB133T forms an independent lineage within the genus Mucilaginibacter. Strain WPCB133T was distantly related to Mucilaginibacter kameinonensis SCKT (94.7 % sequence similarity), Mucilaginibacter paludis TPT56T (94.5 %) and Mucilaginibacter gracilis TPT18T (94.4 %). Phenotypic characteristics distinguished strain WPCB133T from members of the genus Mucilaginibacter. On the basis of evidence presented in this study, strain WPCB133T represents a novel species of the genus Mucilaginibacter, for which the name Mucilaginibacter rigui sp. nov. is proposed. The type strain is WPCB133T (=KCTC 12534T =NBRC 101115T). An emended description of the genus Mucilaginibacter is also proposed.
-
-
-
Algoriphagus lutimaris sp. nov., isolated from a tidal flat sediment
More LessA Gram-negative, non-motile, non-spore-forming bacterial strain, S1-3T, was isolated from a tidal flat sediment on the west coast of Korea and its taxonomic position was investigated. Strain S1-3T grew optimally at 30 °C and in the presence of 2 % (w/v) NaCl. Strain S1-3T contained MK-7 as the predominant menaquinone and C16 : 1 ω7c and/or iso-C15 : 0 2-OH and iso-C15 : 0 as the major fatty acids. The DNA G+C content was 41.4 mol%. Phylogenetic analyses based on 16S rRNA gene sequences revealed that strain S1-3T fell within the clade comprising Algoriphagus species, clustering with Algoriphagus halophilus IMSNU 14013T, with which it exhibited 99.6 % 16S rRNA gene sequence similarity. The 16S rRNA gene sequence similarity between strain S1-3T and the type strains of other Algoriphagus species was 94.0–97.1 %. Differential phenotypic properties and phylogenetic and genetic distinctiveness of strain S1-3T demonstrated that this strain is distinguishable from the other Algoriphagus species as well as A. halophilus. On the basis of phenotypic, chemotaxonomic, phylogenetic and genetic data, strain S1-3T is considered to represent a novel species of the genus Algoriphagus, for which the name Algoriphagus lutimaris sp. nov. is proposed. The type strain is S1-3T (=KCTC 22630T =CCUG 57608T).
-
-
-
Maribacter stanieri sp. nov., a marine bacterium of the family Flavobacteriaceae
More LessThe taxonomic status of two novel heterotrophic, Gram-negative, gliding and yellow pigmented bacterial strains was established in this study. Phylogenetic analysis based on 16S rRNA gene sequencing revealed that the strains formed a distinct lineage within the genus Maribacter, a member of the family Flavobacteriaceae, with sequence similarities of 96.3–98.5 % to recognized species of the genus Maribacter. The maximum growth temperature of the strains was 35 °C and they required NaCl or seawater for growth. They hydrolysed aesculin and gelatin, reduced nitrates to nitrites and produced acid from carbohydrates. The DNA G+C contents of strains KMM 6025 and KMM 6046T were 36–37 mol%. On the basis of phenotypic, genotypic and phylogenetic characteristics, it is suggested that the new isolates represent a novel species of the genus Maribacter, for which the name Maribacter stanieri sp. nov. is proposed. The type strain is KMM 6046T (=KCTC 22023T=LMG 22581T).
-
-
-
Pedobacter glucosidilyticus sp. nov., isolated from dry riverbed soil
More LessTwo Gram-staining-negative, rod-shaped, non-spore-forming bacterial strains, 1-2T and 1-4 were isolated from dry riverbed soil collected from the Xietongmen area of Tibet, China. On the basis of 16S rRNA gene sequence similarity, the novel strains were shown to belong to the genus Pedobacter, sharing <95 % sequence similarity with all recognized species of the genus Pedobacter. The major respiratory quinone was MK-7 and the predominant cellular fatty acids were iso-C15 : 0, iso-C17 : 0 3-OH and summed feature 3 (comprising iso-C16 : 1 ω7c and/or C16 : 1 ω6c). The DNA G+C contents were 37.2–37.6 mol%. Chemotaxonomic data supported the affiliation of the two new isolates to the genus Pedobacter and the results of physiological and biochemical tests confirmed that the new strains differed significantly from the recognized species of the genus Pedobacter. Therefore, the new isolates represent a novel species within the genus Pedobacter, for which the name Pedobacter glucosidilyticus sp. nov. is proposed. The type strain is 1-2T (=CCTCC AB 206110T=KCTC 22438T).
-
- Firmicutes And Related Organisms
-
-
Transfer of Bacillus mucilaginosus and Bacillus edaphicus to the genus Paenibacillus as Paenibacillus mucilaginosus comb. nov. and Paenibacillus edaphicus comb. nov.
More LessBacillus mucilaginosus and Bacillus edaphicus were reclassified based on their 16S rRNA and gyrB gene sequences, DNA–DNA hybridization, fatty acid methyl esters and other taxonomic characteristics. Phylogenetic analysis based on 16S rRNA and gyrB gene sequences indicated that strains of B. mucilaginosus and B. edaphicus were members of the genus Paenibacillus, with over 90.4 % and 70.3 % sequence similarity, respectively. Their DNA G+C contents were 54.5–56.8 mol%. The DNA–DNA relatedness values of B. edaphicus VKPM B-7517T with B. mucilaginosus KNP414 and B. mucilaginosus CGMCC 1.236 were 89.2 % and 88.7 %, respectively. The major isoprenoid quinone of B. mucilaginosus and B. edaphicus was MK-7 (94.1–95.7 %). The peptidoglycan type was A1γ (meso-diaminopimelic acid) and the major polar lipids were phosphatidylglycerol and diphosphatidylglycerol. The major fatty acids were anteiso-C15 : 0, C16 : 1 ω11c and C16 : 0. Phenotypic features and fatty acid profiles supported the similarity of B. mucilaginosus and B. edaphicus to Paenibacillus validus CCTCC 95016T and confirmed their relationship with members of the genus Paenibacillus. Therefore, it is proposed that Bacillus mucilaginosus and Bacillus edaphicus be transferred to the genus Paenibacillus as Paenibacillus mucilaginosus comb. nov. (type strain HSCC 1605T=VKPM B-7519T=VKM B-1480DT=CIP 105815T=KCTC 3870T) and Paenibacillus edaphicus comb. nov. (type strain VKPM B-7517T=DSM 12974T=CIP 105814T), respectively.
-
-
-
Jeotgalibacillus salarius sp. nov., isolated from a marine saltern, and reclassification of Marinibacillus marinus and Marinibacillus campisalis as Jeotgalibacillus marinus comb. nov. and Jeotgalibacillus campisalis comb. nov., respectively
More LessA Gram-variable, motile and rod-shaped bacterial strain, ASL-1T, was isolated from a marine saltern located on the coast of the Yellow Sea, Korea. A neighbour-joining phylogenetic tree based on 16S rRNA gene sequences showed that strain ASL-1T clustered with Jeotgalibacillus alimentarius YKJ-13T and that this cluster joined the clade comprising the type strains of two Marinibacillus species. Strain ASL-1T exhibited 16S rRNA gene sequence similarity values of 97.3 % to J. alimentarius YKJ-13T and 96.5 % to the type strains of Marinibacillus marinus and Marinibacillus campisalis. The chemotaxonomic properties of strain ASL-1T were similar to those of one or two of the genera Jeotgalibacillus and Marinibacillus. The peptidoglycan type was A1α linked directly through l-lysine as the diamino acid. Strain ASL-1T contained MK-7 as the predominant menaquinone with the presence of a significant amount of MK-8. The predominant fatty acid was anteiso-C15 : 0. The DNA G+C content was 42.9 mol%. Differential phenotypic properties, together with the phylogenetic and genetic distinctiveness, revealed that strain ASL-1T could be differentiated from J. alimentarius and the two Marinibacillus species. On the basis of the data presented, strain ASL-1T represents a novel species within the genus Jeotgalibacillus, for which the name Jeotgalibacillus salarius sp. nov. is proposed. The type strain is ASL-1T (=KCTC 13257T=CCUG 56751T). It is also proposed that Marinibacillus marinus and Marinibacillus campisalis be reclassified as Jeotgalibacillus marinus comb. nov. (type strain 581T=DSM 1297T=ATCC 29841T=CCUG 28884T=CIP 103308T=LMG 6930T) and Jeotgalibacillus campisalis comb. nov. (type strain SF-57T=KCCM 41644T=JCM 11810T), respectively.
-
-
-
Bacillus trypoxylicola sp. nov., xylanase-producing alkaliphilic bacteria isolated from the guts of Japanese horned beetle larvae (Trypoxylus dichotomus septentrionalis)
More LessThree xylanase-producing alkaliphilic strains, SU1T, 36AC4 and 36AC6, were isolated from the guts of larvae of the Japanese horned beetle (Trypoxylus dichotomus septentrionalis). The isolates stained Gram-positive and were aerobic, spore-forming, non-motile and rod-shaped and grew optimally at 30 °C and pH 9. They contained MK-7 as the major isoprenoid quinone and iso-C15 : 0, anteiso-C15 : 0, anteiso-C17 : 0 and iso-C17 : 0 as the major fatty acids. The DNA G+C contents of the strains were 37.4–37.7 mol%. On the basis of 16S rRNA gene sequence similarity, these strains were shown to belong to the genus Bacillus. Although their 16S rRNA gene sequence similarity to the type strains of the alkaliphilic species Bacillus pseudalcaliphilus and B. alcalophilus was 97 %, the novel isolates formed a distinct group in the phylogenetic trees and DNA–DNA relatedness values to the type strains of these species were less than 30 %. Results of physiological and biochemical tests, including salt preference, enabled these strains to be differentiated phenotypically from described Bacillus species. Therefore, strains SU1T, 36AC4 and 36AC6 represent a novel species for which the name Bacillus trypoxylicola sp. nov. is proposed; the type strain is SU1T (=NBRC 102646T =KCTC 13244T).
-
-
-
Caldicoprobacter oshimai gen. nov., sp. nov., an anaerobic, xylanolytic, extremely thermophilic bacterium isolated from sheep faeces, and proposal of Caldicoprobacteraceae fam. nov.
More LessAn obligately anaerobic, xylanolytic, extremely thermophilic bacterium, strain JW/HY-331T, was isolated from sheep faeces collected from a farm at the University of Georgia, USA. Cells of strain JW/HY-331T stained Gram-positive and were catalase-negative, non-motile rods. Single terminal endospores (0.4–0.6 μm in diameter) swelled the mother cell. Growth ranges were 44–77 °C (optimum 70 °C at pH70 °C 7.2) and pH70 °C 5.9–8.6 (optimum 7.2 at 70 °C). Salt tolerance was 0–2.0 % (w/v) NaCl. No growth was observed at or below 42 °C or at or above 79 °C or at pH70 °C 5.7 and below or 8.9 and above. In the presence of 0.3 % yeast extract and 0.1 % tryptone, strain JW/HY-331T utilized xylose, glucose, galactose, cellobiose, raffinose and xylan as carbon and energy sources, but not dextran, soluble potato starch, CM-cellulose, cellulose powder, casein or Casamino acids. Fermentation products from glucose were lactate, acetate, ethanol, CO2 and H2. The G+C content of the genomic DNA was 45.4 mol% (HPLC). Major cellular fatty acids were iso-C17 : 0, iso-C15 : 0 and anteiso-C17 : 0. No respiratory quinones were detected. The cell-wall structure was a single layer (Gram-type positive) of the peptidoglycan type A1γ; the cell-wall sugars were galactose and mannose. Based on 16S rRNA gene sequence analysis, ‘Catabacter hongkongensis’ HKU16 (85.4 % similarity), Caloramator fervidus ATCC 43204T (84.2 %) and Caloranaerobacter azorensis MV1087T (83.4 %) were the closest relatives, but they were only distantly related to strain JW/HY-331T. On the basis of physiological, chemotaxonomic and phylogenetic data, isolate JW/HY-331T (=DSM 21659T =ATCC BAA-1711T) is proposed as the type strain of Caldicoprobacter oshimai gen. nov., sp. nov., placed in Caldicoprobacteraceae fam. nov. within the order Clostridiales of the phylum Firmicutes.
-
-
-
Streptococcus porci sp. nov., isolated from swine sources
More LessTwo unidentified Gram-positive, catalase-negative, coccus-shaped organisms were recovered from pigs and subjected to a polyphasic taxonomic analysis. Based on cellular morphology and biochemical criteria, the isolates were tentatively assigned to the genus Streptococcus, although the organisms did not appear to correspond to any recognized species. Comparative 16S rRNA gene sequence studies confirmed this identification and showed that the nearest phylogenetic relatives of the unknown cocci were Streptococcus plurextorum 1956-02T and Streptococcus suis NCTC 10234T (97.9 and 96.0 % 16S rRNA gene sequence similarity, respectively). The new isolates were related most closely to S. suis CIP 103217T based on rpoB gene sequence analysis (<8 % sequence divergence). DNA–DNA pairing studies showed that one of the unidentified strains (2923-03T) displayed DNA relatedness values of 26.6 and 27.2 % with S. plurextorum CECT 7308T and S. suis NCTC 10234T, respectively. On the basis of phenotypic and phylogenetic evidence, it is proposed that the unknown isolates from pigs be classified in the genus Streptococcus as members of Streptococcus porci sp. nov., with the type strain 2923-03T (=CECT 7374T =CCUG 55896T).
-
-
-
Lactobacillus equicursoris sp. nov., isolated from the faeces of a thoroughbred racehorse
More LessWe previously isolated five strains of putative lactobacilli from the faeces of a thoroughbred horse (a 4-year-old male). Of the five strains, four were identified as members of existing Lactobacillus species; however, sequence analysis of the 16S rRNA gene revealed that the fifth isolate, DI70T, showed approximately 97 % identity (1325/1366 bp) with the type strain of Lactobacillus delbrueckii. Therefore, we considered the possibility that DI70T represents a novel species of the genus Lactobacillus. Cells of strain DI70T were Gram-stain-positive, catalase-negative, non-spore-forming, non-motile rods. In phylogenetic trees constructed on the basis of 16S rRNA gene sequences, strain DI70T formed a subcluster in the L. delbrueckii phylogenetic group and was closely related to L. delbrueckii, Lactobacillus crispatus and Lactobacillus jensenii. However, analysis of DNA–DNA relatedness showed that DI70T was genetically distinct from its phylogenetic relatives. The isolate also exhibited distinct biochemical and physiological characteristics when compared with its phylogenetic relatives. It required anaerobic conditions for growth on agar medium. The results indicate that isolate DI70T indeed represents a novel species of the genus Lactobacillus, for which we propose the name Lactobacillus equicursoris sp. nov. The type strain is DI70T (=JCM 14600T =DSM 19284T).
-
-
-
Paenibacillus riograndensis sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Triticum aestivum
More LessA bacterial strain designated SBR5T was isolated from the rhizosphere of Triticum aestivum. A phylogenetic analysis based on the 16S rRNA gene sequence placed the isolate within the genus Paenibacillus, being most closely related to Paenibacillus graminis RSA19T (98.1 % similarity). The isolate was a Gram-reaction-variable, motile, facultatively anaerobic bacterium, with spores in a terminal position in cells. Starch was utilized and dihydroxyacetone and catalase were produced. Strain SBR5T displayed plant-growth-promoting rhizobacteria characteristics: the ability to fix nitrogen and to produce siderophores and indole-3-acetic acid. The DNA G+C content was 55.1 mol%. Chemotaxonomic analysis of the isolated strain revealed that MK-7 was the predominant menaquinone, while the major fatty acid was anteiso-C15 : 0. DNA–DNA hybridization values between strain SBR5T and P. graminis RSA19T, Paenibacillus odorifer TOD45T and Paenibacillus borealis KK19T were 43, 35 and 28 %, respectively. These DNA relatedness data and the results of phylogenetic and phenotypic analyses showed that strain SBR5T should be considered as the nitrogen-fixing type strain of a novel species of the genus Paenibacillus, for which the name Paenibacillus riograndensis sp. nov. is proposed. The type strain is SBR5T (=CCGB 1313T =CECT 7330T).
-
-
-
Lactobacillus similis sp. nov., isolated from fermented cane molasses
More LessThe taxonomic position of strain JCM 2765T isolated from fermented cane molasses in Thailand was reinvestigated. Strain JCM 2765T was originally identified as representing Lactobacillus buchneri on the basis of biochemical and physiological characteristics. In the present study, 16S rRNA gene sequence analysis of strain JCM 2765T demonstrated a low level of similarity with the type strain of L. buchneri (92.5 %) and high levels with those of Lactobacillus collinoides (97.6 %) and Lactobacillus paracollinoides (98.0 %). Ribotyping was applied to investigate the relationships between strain JCM 2765T, L. collinoides and L. paracollinoides. The dendrogram based on ribotyping patterns showed one cluster for six strains of L. paracollinoides, and that strain JCM 2765T and L. collinoides JCM 1123T were each independent. Based on additional phenotypic findings and DNA–DNA hybridization results, strain JCM 2765T is considered to represent a novel species of the genus Lactobacillus, for which the name Lactobacillus similis sp. nov. is proposed. The type strain is JCM 2765T (=LMG 23904T).
-
- Other Bacteria
-
-
Thermodesulfatator atlanticus sp. nov., a thermophilic, chemolithoautotrophic, sulfate-reducing bacterium isolated from a Mid-Atlantic Ridge hydrothermal vent
More LessA novel, strictly anaerobic, thermophilic, sulfate-reducing bacterium, designated strain AT1325T, was isolated from a deep-sea hydrothermal vent at the Rainbow site on the Mid-Atlantic Ridge. This strain was subjected to a polyphasic taxonomic analysis. Cells were Gram-negative motile rods (approximately 2.4×0.6 μm) with a single polar flagellum. Strain AT1325T grew at 55–75 °C (optimum, 65–70 °C), at pH 5.5–8.0 (optimum, 6.5–7.5) and in the presence of 1.5–4.5 % (w/v) NaCl (optimum, 2.5 %). Cells grew chemolithoautotrophically with H2 as an energy source and
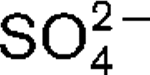 as an electron acceptor. Alternatively, the novel isolate was able to use methylamine, peptone or yeast extract as carbon sources. The dominant fatty acids (>5 % of the total) were C16 : 0, C18 : 1
ω7c, C18 : 0 and C19 : 0 cyclo ω8c. The G+C content of the genomic DNA of strain AT1325T was 45.6 mol%. Phylogenetic analyses based on 16S rRNA gene sequences placed strain AT1325T within the family Thermodesulfobacteriaceae, in the bacterial domain. Comparative 16S rRNA gene sequence analysis indicated that strain AT1325T belonged to the genus Thermodesulfatator, sharing 97.8 % similarity with the type strain of Thermodesulfatator indicus, the unique representative species of this genus. On the basis of the data presented, it is suggested that strain AT1325T represents a novel species of the genus Thermodesulfatator, for which the name Thermodesulfatator atlanticus sp. nov. is proposed. The type strain is AT1325T (=DSM 21156T=JCM 15391T).
as an electron acceptor. Alternatively, the novel isolate was able to use methylamine, peptone or yeast extract as carbon sources. The dominant fatty acids (>5 % of the total) were C16 : 0, C18 : 1
ω7c, C18 : 0 and C19 : 0 cyclo ω8c. The G+C content of the genomic DNA of strain AT1325T was 45.6 mol%. Phylogenetic analyses based on 16S rRNA gene sequences placed strain AT1325T within the family Thermodesulfobacteriaceae, in the bacterial domain. Comparative 16S rRNA gene sequence analysis indicated that strain AT1325T belonged to the genus Thermodesulfatator, sharing 97.8 % similarity with the type strain of Thermodesulfatator indicus, the unique representative species of this genus. On the basis of the data presented, it is suggested that strain AT1325T represents a novel species of the genus Thermodesulfatator, for which the name Thermodesulfatator atlanticus sp. nov. is proposed. The type strain is AT1325T (=DSM 21156T=JCM 15391T).
-
- Proteobacteria
-
-
Pandoraea thiooxydans sp. nov., a facultatively chemolithotrophic, thiosulfate-oxidizing bacterium isolated from rhizosphere soils of sesame (Sesamum indicum L.)
More LessA facultatively chemolithoautotrophic, thiosulfate-oxidizing, Gram-negative, aerobic, motile, rod-shaped bacterial strain, designated ATSB16T, was isolated from rhizosphere soils of sesame (Sesamum indicum L.). 16S rRNA gene sequence analysis demonstrated that this strain was closely related to Pandoraea pnomenusa LMG 18087T (96.7 % similarity), P. pulmonicola LMG 18016T (96.5 %), P. apista LMG 16407T (96.2 %), P. norimbergensis LMG 18379T (96.1 %) and P. sputorum LMG 18819T (96.0 %). Strain ATSB16T shared 96.0–96.4 % sequence similarity with four unnamed genomospecies of Pandoraea. The major cellular fatty acids of the strain ATSB16T were C17 : 0 cyclo (33.0 %) and C16 : 0 (30.6 %). Q-8 was the predominant respiratory quinone. The major polar lipids were phosphatidylmethylethanolamine, diphosphatidylglycerol, phosphatidylethanolamine and two unidentified aminophospholipids. Hydroxyputrescine and putrescine were the predominant polyamines. The genomic DNA G+C content of the strain was 64.0 mol%. On the basis of the results obtained from this study, strain ATSB16T represents a novel species of the genus Pandoraea, for which the name Pandoraea thiooxydans sp. nov. is proposed. The type strain is ATSB16T (=KACC 12757T =LMG 24779T).
-
-
-
Basfia succiniciproducens gen. nov., sp. nov., a new member of the family Pasteurellaceae isolated from bovine rumen
More LessGram-negative, coccoid, non-motile bacteria that are catalase-, urease- and indole-negative, facultatively anaerobic and oxidase-positive were isolated from the bovine rumen using an improved selective medium for members of the Pasteurellaceae. All strains produced significant amounts of succinic acid under anaerobic conditions with glucose as substrate. Phenotypic characterization and multilocus sequence analysis (MLSA) using 16S rRNA, rpoB, infB and recN genes were performed on seven independent isolates. All four genes showed high sequence similarity to their counterparts in the genome sequence of the patent strain MBEL55E, but less than 95 % 16S rRNA gene sequence similarity to any other species of the Pasteurellaceae. Genetically these strains form a very homogeneous group in individual as well as combined phylogenetic trees, clearly separated from other genera of the family from which they can also be separated based on phenotypic markers. Genome relatedness as deduced from the recN gene showed high interspecies similarities, but again low similarity to any of the established genera of the family. No toxicity towards bovine, human or fish cells was observed and no RTX toxin genes were detected in members of the new taxon. Based on phylogenetic clustering in the MLSA analysis, the low genetic similarity to other genera and the phenotypic distinction, we suggest to classify these bovine rumen isolates as Basfia succiniciproducens gen. nov., sp. nov. The type strain is JF4016T (=DSM 22022T =CCUG 57335T).
-
-
-
Salinarimonas rosea gen. nov., sp. nov., a new member of the α-2 subgroup of the Proteobacteria
More LessA Gram-negative, rod-shaped, facultatively anaerobic, halotolerant bacterial strain, designated YIM YD3T, was isolated from a salt mine in Yunnan, south-west China. The taxonomy of strain YIM YD3T was investigated by a polyphasic approach. Strain YIM YD3T was motile, formed pink colonies and was positive for catalase and oxidase activities. Q-10 was the predominant respiratory ubiquinone. The major polar lipids were diphosphatidylglycerol, phosphatidylglycerol, phosphatidylmethylethanolamine, phosphatidylcholine and two unknown phospholipids. The major fatty acids (>10 % of total fatty acids) were C18 : 1 ω7c, C18 : 1 ω9c, C16 : 0 and C19 : 0 cyclo ω8c. The DNA G+C content was 71.8 mol%. Phylogenetic analysis based on 16S rRNA gene sequence comparisons showed that the isolate formed a distinct line within a clade containing the genera Balneimonas, Bosea, Chelatococcus and Microvirga in the order Rhizobiales, with highest levels of 16S RNA gene sequence similarity to the type strain of Balneimonas flocculans (93.5 %). On the basis of phenotypic, chemotaxonomic and phylogenetic data, strain YIM YD3T represents a novel species in a new genus, for which the name Salinarimonas rosea gen. nov., sp. nov. is proposed, with strain YIM YD3T (=KCTC 22346T=CCTCC AA208038T) as the type strain.
-
-
-
Aeromonas fluvialis sp. nov., isolated from a Spanish river
More LessA Gram-stain-negative, facultatively anaerobic bacterial strain, designated 717T, was isolated from a water sample collected from the Muga river, Girona, north-east Spain. Preliminary analysis of the 16S rRNA gene sequence showed that this strain belonged to the genus Aeromonas, the nearest species being Aeromonas veronii (99.5 % similarity, with seven different nucleotides). A polyphasic study based on a multilocus phylogenetic analysis of five housekeeping genes (gyrB, rpoD, recA, dnaJ and gyrA; 3684 bp) showed isolate 717T to be an independent phylogenetic line, with Aeromonas sobria, Aeromonas veronii and Aeromonas allosaccharophila as the closest neighbour species. DNA–DNA reassociation experiments and phenotypic analysis identified that strain 717T represents a novel species, for which the name Aeromonas fluvialis sp. nov. is proposed, with type strain 717T (=CECT 7401T =LMG 24681T).
-
-
-
Psychromonas boydii sp. nov., a gas-vacuolate, psychrophilic bacterium isolated from an Arctic sea-ice core
More LessA gas-vacuolate bacterium, strain 174T, was isolated from a sea-ice core collected from Point Barrow, Alaska, USA. Comparative analysis of 16S rRNA gene sequences showed that this bacterium was most closely related to Psychromonas ingrahamii 37T, with a similarity of >99 %. However, strain 174T could be clearly distinguished from closely related species by DNA–DNA hybridization; relatedness values determined by two different methods between strain 174T and P. ingrahamii 37T were 58.4 and 55.7 % and those between strain 174T and Psychromonas antarctica DSM 10704T were 46.1 and 33.1 %, which are well below the 70 % level used to define a distinct species. Phenotypic analysis, including cell size (strain 174T is the largest member of the genus Psychromonas, with rod-shaped cells, 8–18 μm long), further differentiated strain 174T from other members of the genus Psychromonas. Strain 174T could be distinguished from its closest relative, P. ingrahamii, by its utilization of d-mannose and d-xylose as sole carbon sources, its ability to ferment myo-inositol and its inability to use fumarate and glycerol as sole carbon sources. In addition, strain 174T contained gas vacuoles of two distinct morphologies and grew at temperatures ranging from below 0 to 10 °C and its optimal NaCl concentration for growth was 3.5 %. The DNA G+C content was 40 mol%. Whole-cell fatty acid analysis showed that 16 : 1ω7c and 16 : 0 comprised 44.9 and 26.4 % of the total fatty acid content, respectively. The name Psychromonas boydii sp. nov. is proposed for this novel species, with strain 174T (=DSM 17665T =CCM 7498T) as the type strain.
-
-
-
Taxonomic study of Marinomonas strains isolated from the seagrass Posidonia oceanica, with descriptions of Marinomonas balearica sp. nov. and Marinomonas pollencensis sp. nov.
More LessNovel aerobic, Gram-negative bacteria with DNA G+C contents below 50 mol% were isolated from the culturable microbiota associated with the Mediterranean seagrass Posidonia oceanica. 16S rRNA gene sequence analyses revealed that they belong to the genus Marinomonas. Strain IVIA-Po-186 is a strain of the species Marinomonas mediterranea, showing 99.77 % 16S rRNA gene sequence similarity with the type strain, MMB-1T, and sharing all phenotypic characteristics studied. This is the first description of this species forming part of the microbiota of a marine plant. A second strain, designated IVIA-Po-101T, was closely related to M. mediterranea based on phylogenetic studies. However, it differed in characteristics such as melanin synthesis and tyrosinase, laccase and antimicrobial activities. In addition, strain IVIA-Po-101T was auxotrophic and unable to use acetate. IVIA-Po-101T shared 97.86 % 16S rRNA gene sequence similarity with M. mediterranea MMB-1T, but the level of DNA–DNA relatedness between the two strains was only 10.3 %. On the basis of these data, strain IVIA-Po-101T is considered to represent a novel species of the genus Marinomonas, for which the name Marinomonas balearica sp. nov. is proposed. The type strain is IVIA-Po-101T (=CECT 7378T =NCIMB 14432T). A third novel strain, IVIA-Po-185T, was phylogenetically distant from all recognized Marinomonas species. It shared the highest 16S rRNA gene sequence similarity (97.4 %) with the type strain of Marinomonas pontica, but the level of DNA–DNA relatedness between the two strains was only 14.5 %. A differential chemotaxonomic marker of this strain in the genus Marinomonas is the presence of the fatty acid C17 : 0 cyclo. Strain IVIA-Po-185T is thus considered to represent a second novel species of the genus, for which the name Marinomonas pollencensis sp. nov. is proposed. The type strain is IVIA-Po-185T (=CECT 7375T =NCIMB 14435T). An emended description of the genus Marinomonas is given based on the description of these two novel species, as well as other Marinomonas species described after the original description of the genus.
-
-
-
Shinella fusca sp. nov., isolated from domestic waste compost
More LessA bacterium, designated strain DC-196T, isolated from kitchen refuse compost was analysed by using a polyphasic approach. Strain DC-196T was characterized as a Gram-negative short rod that was catalase- and oxidase-positive, and able to grow at 10–40 °C, pH 6–9 and in NaCl concentrations as high as 3 %. Chemotaxonomically, C18 : 1 was observed to be the predominant cellular fatty acid and ubiquinone 10 (Q10) was the predominant respiratory quinone. The G+C content of the genomic DNA was determined to be 66 mol%. On the basis of the genotypic, phenotypic and chemotaxonomic characteristics, strain DC-196T was assigned to the genus Shinella, although with distinctive features. At the time of writing, 16S rRNA gene sequence similarities of 97.6–96.8 % and the low DNA–DNA hybridization values of 38.2–32.2 % with the type strains of the three recognized Shinella species confirmed that strain DC-196T represents a novel species of the genus, for which the name Shinella fusca sp. nov. is proposed (type strain DC-196T=CCUG 55808T=LMG 24714T).
-
-
-
Polynucleobacter cosmopolitanus sp. nov., free-living planktonic bacteria inhabiting freshwater lakes and rivers
More LessFive heterotrophic, aerobic, catalase- and oxidase-positive, non-motile strains were characterized from freshwater habitats located in Austria, France, Uganda, P. R. China and New Zealand. The strains shared 16S rRNA gene similarities of ≥99.3 %. The novel strains grew on NSY medium over a temperature range of 10–35 °C (two strains also grew at 5 °C and one strain grew at 38 °C) and a NaCl tolerance range of 0.0–0.3 % (four strains grew up to 0.5 % NaCl). The predominant fatty acids were C16 : 0, C18 : 1 ω7c, C12 : 0 3-OH, and summed feature 3 (including C16 : 1 ω7c). The DNA G+C content of strain MWH-MoIso2T was 44.9 mol%. Phylogenetic analysis of 16S rRNA gene sequences demonstrated that the five new strains formed a monophyletic cluster closely related to Polynucleobacter necessarius (96–97 % sequence similarity). This cluster also harboured other isolates as well as environmental sequences which have been obtained from several habitats. Investigations with taxon-specific FISH probes demonstrated that the novel bacteria dwell as free-living, planktonic cells in freshwater systems. Based on the revealed phylogeny and pronounced chemotaxonomic differences to P. necessarius (presence of >7 % C12 : 0 3-OH and absence of C12 : 0 and C12 : 0 2-OH), the new strains are suggested to represent a novel species, for which the name Polynucleobacter cosmopolitanus sp. nov. is proposed. The type strain is MWH-MoIso2T (=DSM 21490T=CIP 109840T=LMG 25212T). The novel species belongs to the minority of described species of free-living bacteria for which both in situ data from their natural environments and culture-based knowledge are available.
-
-
-
Silvimonas iriomotensis sp. nov. and Silvimonas amylolytica sp. nov., new members of the class Betaproteobacteria isolated from the subtropical zone in Japan
More LessThe taxonomic positions of two bacterial strains, ir6-1T and ir6-4T, isolated from soil collected in Iriomote Island in Japan, were determined by using a polyphasic taxonomic approach. The strains were facultatively anaerobic, motile and Gram-stain-negative rods and their optimum pH for growth was pH 4.0. Their major respiratory quinone was ubiquinone-8 and the predominant cellular fatty acids were C14 : 0, C16 : 0, C16 : 1 ω7c and C18 : 1 ω7c. The G+C content of the genomic DNA of ir6-1T and ir6-4T was 59.9 and 57.5 mol%, respectively. Phylogenetic analysis based on 16S rRNA gene sequences showed that the strains clustered with the genus Silvimonas in the class Betaproteobacteria. DNA–DNA similarities were lower than 53 % among ir6-1T, ir6-4T and Silvimonas terrae NBRC 100961T, and these strains could be differentiated from each other by several phenotypic characters. Based on these results, we propose the emendation of the genus Silvimonas and inclusion of two novel species, Silvimonas iriomotensis sp. nov. (type strain ir6-1T=NBRC 103188T =CGMCC 1.8859T =KCTC 22513T) and Silvimonas amylolytica sp. nov. (ir6-4T =NBRC 103189T =CGMCC 1.8860T =KCTC 22514T).
-
-
-
Mangrovibacter plantisponsor gen. nov., sp. nov., a nitrogen-fixing bacterium isolated from a mangrove-associated wild rice (Porteresia coarctata Tateoka)
More LessA facultatively anaerobic, nitrogen-fixing bacterium, strain MSSRF40T, was isolated from roots of mangrove-associated wild rice (Porteresia coarctata Tateoka). On the basis of 16S rRNA gene sequence similarities, strain MSSRF40T was shown to belong to the family Enterobacteriaceae, most closely related to Cronobacter muytjensii E603T (97.2 % sequence similarity), Enterobacter cloacae subsp. dissolvens LMG 2683T (97.1 %), E. radicincitans D5/23T (97.1 %) and E. ludwigii EN-119T (97.0 %). Sequence analysis of rpoB, gyrB and hsp60 genes showed that strain MSSRF40T had relatively low sequence similarity (<91, <84 and <90 %) to recognized species of different genera of the family Enterobacteriaceae and formed an independent phyletic lineage in all phylogenetic analyses using the 16S rRNA, rpoB, gyrB and hsp60 genes, clearly indicating that strain MSSRF40T could not be affiliated to any of the recognized genera within the family Enterobacteriaceae. The dominant cellular fatty acids were C16 : 0, C16 : 1 ω7c and/or iso-C15 : 0 2-OH and C18 : 1 ω7c, similar to those of other members of the Enterobacteriaceae. The DNA G+C content was 50.1 mol%. Phylogenetic distinctiveness and phenotypic differences from its phylogenetic neighbours indicated that strain MSSRF40T represents a novel species and genus within the family Enterobacteriaceae, for which the name Mangrovibacter plantisponsor gen. nov., sp. nov. is proposed. The type strain of Mangrovibacter plantisponsor is strain MSSRF40T (=LMG 24236T =DSM 19579T).
-
-
-
Jannaschia seohaensis sp. nov., isolated from a tidal flat sediment
More LessA Gram-negative, motile and pleomorphic bacterial strain, SMK-146T, was isolated from a tidal flat sediment of the Yellow Sea, Korea, and its taxonomic position was investigated. Strain SMK-146T grew optimally at pH 7.0–8.0 and 30 °C. It contained Q-10 as the predominant ubiquinone and C18 : 1 ω7c and 11-methyl C18 : 1 ω7c as the major fatty acids. The major polar lipids were phosphatidylcholine, phosphatidylglycerol and phosphatidylethanolamine. The DNA G+C content was 68.4 mol%. Phylogenetic analysis based on 16S rRNA gene sequences showed that strain SMK-146T belongs to the genus Jannaschia. Strain SMK-146T exhibited 16S rRNA gene sequence similarity values of 95.3–97.0 % to the type strains of the five recognized Jannaschia species. The mean DNA–DNA relatedness value between strain SMK-146T and Jannaschia seosinensis KCCM 42114T, the closest phylogenetic neighbour, was 17 %. Differential phenotypic properties also revealed that strain SMK-146T differs from the recognized Jannaschia species. On the basis of phenotypic, phylogenetic and genetic data, strain SMK-146T represents a novel species of the genus Jannaschia, for which the name Jannaschia seohaensis sp. nov. is proposed. The type strain is SMK-146T (=KCTC 22172T =CCUG 55326T).
-
-
-
Gaetbulicola byunsanensis gen. nov., sp. nov., isolated from tidal flat sediment
More LessA Gram-negative, non-motile and pleomorphic bacterial strain, SMK-114T, which belongs to the class Alphaproteobacteria, was isolated from a tidal flat sample collected in Byunsan, Korea. Strain SMK-114T grew optimally at pH 7.0–8.0 and 25–30 °C and in the presence of 2 % (w/v) NaCl. A neighbour-joining phylogenetic tree based on 16S rRNA gene sequences showed that strain SMK-114T formed a cluster with Octadecabacter species, with which it exhibited 16S rRNA gene sequence similarity values of 95.2–95.4 %. This cluster was part of the clade comprising Thalassobius species with a bootstrap resampling value of 76.3 %. Strain SMK-114T exhibited 16S rRNA gene sequence similarity values of 95.1–96.3 % to members of the genus Thalassobius. It contained Q-10 as the predominant ubiquinone and C18 : 1 ω7c as the major fatty acid. The DNA G+C content was 60.0 mol%. On the basis of phenotypic, chemotaxonomic and phylogenetic data, strain SMK-114T is considered to represent a novel species in a new genus for which the name Gaetbulicola byunsanensis gen. nov., sp. nov. is proposed. The type strain of Gaetbulicola byunsanensis is SMK-114T (=KCTC 22632T =CCUG 57612T).
-
-
-
Psychrobacter piscatorii sp. nov., a psychrotolerant bacterium exhibiting high catalase activity isolated from an oxidative environment
More LessA Gram-negative, non-motile, psychrotolerant bacterium exhibiting high catalase activity, designated strain T-3-2T, was isolated from a drain of a fish-processing plant. Its catalase activity was 12 000 U (mg protein)−1, much higher than the activity of the other Psychrobacter strains tested. The strain grew at 0–30 °C and in the presence of 0–12 % NaCl. The predominant isoprenoid quinone was ubiquinone-8 (Q-8), and C16 : 1 ω9c and C18 : 1 ω9c were the predominant cellular fatty acids. The DNA G+C content of strain T-3-2T was 43.9 mol%. 16S rRNA gene sequence phylogeny suggested that strain T-3-2T is a member of the genus Psychrobacter, with the closest relatives being the type strains of Psychrobacter nivimaris (99.2 % similarity), P. aquimaris (98.7 %) and P. proteolyticus (98.5 %). DNA–DNA hybridization showed less than 65 % relatedness with these strains. A phylogenetic tree based on gyrB gene sequences was more reliable, with higher bootstrap values than the 16S rRNA gene sequence-based tree. The result also differentiated the isolate from previously reported Psychrobacter species. Owing to the significant differences in phenotypic and chemotaxonomic characteristics and the phylogenetic and DNA–DNA relatedness data, the isolate merits classification within a novel species, for which the name Psychrobacter piscatorii sp. nov. is proposed. The type strain is T-3-2T (=JCM 15603T =NCIMB 14510T).
-
-
-
Thalassobaculum salexigens sp. nov., a new member of the family Rhodospirillaceae from the NW Mediterranean Sea, and emended description of the genus Thalassobaculum
More LessA novel Gram-negative bacteria, named CZ41_10aT, was isolated from coastal surface waters of the north-western Mediterranean Sea. Cells were motile, pleomorphic rods, 1.6 μm long and 0.7 μm wide and formed cream colonies on marine agar medium. The G+C content of the genomic DNA was 65 mol%. Phylogenetic analysis of 16S rRNA gene sequences placed the new isolate in the genus Thalassobaculum, a member of the family Rhodospirillaceae, class Alphaproteobacteria. Unlike Thalassobaculum litoreum CL-GR58T, its closest relative, strain CZ41_10aT was unable to grow anaerobically and did not exhibit nitrate reductase activity. On the basis of DNA–DNA hybridization, fatty acid content and physiological and biochemical characteristics, this isolate represents a novel species for which the name Thalassobaculum salexigens sp. nov. is proposed. The type strain is CZ41_10aT (=DSM 19539T=CIP 109064T=MOLA 84T). An emended description of the genus Thalassobaculum is also given.
-
-
-
Oceanibaculum pacificum sp. nov., isolated from hydrothermal field sediment of the south-west Pacific Ocean
More LessA taxonomic study was carried out on strain LMC2up-L3T, which was isolated from a polycyclic aromatic hydrocarbon-degrading consortium enriched with a sediment sample collected from a hydrothermal field of the south-west Pacific Ocean. Phylogenetic analysis based on 16S rRNA gene sequences indicated that strain LMC2up-L3T belonged to the genus Oceanibaculum, with the highest sequence similarity of 98.4 % to Oceanibaculum indicum P24T; similarity to other strains was below 93.1 %. DNA–DNA hybridization between strain LMC2up-L3T and O. indicum P24T was 31 %. rep-PCR fingerprints also differentiated strain LMC2up-L3T from O. indicum P24T. The G+C content of the chromosomal DNA was 67.7 mol%. The principal fatty acids were C16 : 1 (17.8 %), C16 : 0 (21.2 %), C18 : 1 ω7c (23.6 %), C18 : 0 (4.1 %), C18 : 1 2-OH (4.5 %) and C19 : 0 cyclo ω8c (17.4 %). The combined genotypic and phenotypic data show that strain LMC2up-L3T represents a novel species of the genus Oceanibaculum, for which the name Oceanibaculum pacificum sp. nov. is proposed, with the type strain LMC2up-L3T (=CCTCC AB 209059T =LMG 24859T =MCCC 1A02656T). An emended description of the genus Oceanibaculum is also provided.
-
-
-
Aliivibrio finisterrensis sp. nov., isolated from Manila clam, Ruditapes philippinarum and emended description of the genus Aliivibrio
More LessFour strains isolated from cultured Manila clam, Ruditapes philippinarum, in the north-western coast of Spain were characterized phenotypically and genotypically. Phylogenetic analyses based on the 16S rRNA gene sequences indicated that these bacteria were closely related to Aliivibrio wodanis, Aliivibrio salmonicida, Aliivibrio fischeri and Aliivibrio logei with sequence similarities between 98.1 and 96.0 %. Phylogenetic analysis based on the RNA polymerase alpha chain (rpoA), RecA protein (recA), the α-subunit of bacterial ATP synthase (atpA) and the uridine monophosphate (UMP) kinase (pyrH) genes and fluorescent amplified fragment length polymorphism experiments clearly showed that these novel isolates form a tight genomic group different from any currently known Aliivibrio species. On the basis of phylogenetic analysis and phenotypic data, the four strains represent a novel taxon, for which the name Aliivibrio finisterrensis sp. nov. is proposed. Several phenotypic features were revealed that discriminate A. finisterrensis from other Aliivibrio species. The type strain is CMJ 11.1T (=CECT 7228T=LMG 23869T).
-
- Eukaryotic Micro-Organisms
-
-
Parabirojimia multinucleata spec. nov. and Anteholosticha scutellum (Cohn, 1866) Berger, 2003, marine ciliates (Ciliophora, Hypotrichida) from tropical waters in southern China, with notes on their small-subunit rRNA gene sequences
More LessFew studies using modern methods have been carried out on ciliated protozoa in tropical marine waters. In the present work, two hypotrichs, Parabirojimia multinucleata spec. nov. and Anteholosticha scutellum (Cohn, 1866) Berger, 2003, collected from Daya Bay in southern China, were investigated morphologically. P. multinucleata is distinguished by the following combination of characters: slender body, without a snout-like protrusion in the frontal field, and about 50 macronuclear nodules. The poorly known A. scutellum has never been investigated using modern methods; hence, a redescription is needed. During the present study, observations of specimens in vivo and following protargol impregnation revealed new information concerning structures such as the cortical granules and the infraciliature. A redescription and improved diagnosis are supplied based on the China population. The small-subunit (SSU) rRNA gene was sequenced for both organisms and comparisons with those of similar congeners clearly support the findings based on morphological studies.
-
-
-
Candida aechmeae sp. nov. and Candida vrieseae sp. nov., novel yeast species isolated from the phylloplane of bromeliads in Southern Brazil
More LessTwo novel yeast species, Candida aechmeae sp. nov. and Candida vrieseae sp. nov., were isolated from bromeliads in Itapuã Park, Rio Grande do Sul, Brazil. These species are genetically isolated from all other currently recognized ascomycetous yeasts based on their sequence divergence in the D1/D2 domain of the LSU rRNA gene. C. aechmeae sp. nov. is phylogenetically close to Candida ubatubensis, a species also isolated from bromeliads in Brazil, but the novel species can be differentiated on the basis of differences in the D1/D2 domain and positive results for the assimilation of l-arabinose, raffinose, inulin and citrate. Candida vrieseae sp. nov. is phylogenetically placed in a clade near Candida membranifaciens that is composed of several species associated with insects, but the novel species can be differentiated from them by the D1/D2 and ITS gene sequences, positive results for the assimilation of nitrite and a negative result for the assimilation of ethylamine. The type strain for Candida aechmeae sp. nov. is BI153T (=CBS 10831T=NRRL Y-48456T) and the type strain for C. vrieseae sp. nov. is BI146T (=CBS 10829T=NRRL Y-48461T).
-
- Methods
-
-
-
Multilocus sequence analysis of the central clade of the genus Vibrio by using the 16S rRNA, recA, pyrH, rpoD, gyrB, rctB and toxR genes
More LessThe central clade of the genus Vibrio, also called the Vibrio core group, comprises six species that are tightly related (DNA–DNA reassociation values are very close to 70 % for most species pairs). Identification of novel strains to the species level within this group is troublesome and results are quite often dependent on the methodology employed. Therefore, this group represents an excellent framework to test the robustness of multilocus sequence analysis (MLSA) not only for inferring phylogeny but also as an identification tool without the need for DNA–DNA hybridization assays. The genes selected, 16S rRNA, recA, pyrH, rpoD, gyrB, rctB and toxR, were amplified by direct PCR from 44 Vibrio core-group strains. Subsequent analysis allowed us to recognize toxR and rpoD as the most resolving individual genes and showed that concatenated sequences of rpoD, rctB and toxR were more useful than concatenated sequences of all seven genes. To validate our conclusions, MLSA similarities have been correlated with DNA–DNA relatedness values obtained in this study and values taken from the literature. Although the seven concatenated genes gave the best correlation, the concatenated sequences of rpoD, rctB and toxR have the practical advantage of showing a considerable gap between the maximal interspecies similarity and the minimal intraspecies similarity recorded, meaning that they can be used quite conveniently for species identification of vibrios.
-
-
- International Committee On Systematics Of Prokaryotes
-
- Taxonomic Note
-
-
Notes on the characterization of prokaryote strains for taxonomic purposes
More LessTaxonomy relies on three key elements: characterization, classification and nomenclature. All three elements are dynamic fields, but each step depends on the one which precedes it. Thus, the nomenclature of a group of organisms depends on the way they are classified, and the classification (among other elements) depends on the information gathered as a result of characterization. While nomenclature is governed by the Bacteriological Code, the classification and characterization of prokaryotes is an area that is not formally regulated and one in which numerous changes have taken place in the last 50 years. The purpose of the present article is to outline the key elements in the way that prokaryotes are characterized, with a view to providing an overview of some of the pitfalls commonly encountered in taxonomic papers.
-
- Retractions
-
Volumes and issues
-
Volume 74 (2024)
-
Volume 73 (2023)
-
Volume 72 (2022 - 2023)
-
Volume 71 (2020 - 2021)
-
Volume 70 (2020)
-
Volume 69 (2019)
-
Volume 68 (2018)
-
Volume 67 (2017)
-
Volume 66 (2016)
-
Volume 65 (2015)
-
Volume 64 (2014)
-
Volume 63 (2013)
-
Volume 62 (2012)
-
Volume 61 (2011)
-
Volume 60 (2010)
-
Volume 59 (2009)
-
Volume 58 (2008)
-
Volume 57 (2007)
-
Volume 56 (2006)
-
Volume 55 (2005)
-
Volume 54 (2004)
-
Volume 53 (2003)
-
Volume 52 (2002)
-
Volume 51 (2001)
-
Volume 50 (2000)
-
Volume 49 (1999)
-
Volume 48 (1998)
-
Volume 47 (1997)
-
Volume 46 (1996)
-
Volume 45 (1995)
-
Volume 44 (1994)
-
Volume 43 (1993)
-
Volume 42 (1992)
-
Volume 41 (1991)
-
Volume 40 (1990)
-
Volume 39 (1989)
-
Volume 38 (1988)
-
Volume 37 (1987)
-
Volume 36 (1986)
-
Volume 35 (1985)
-
Volume 34 (1984)
-
Volume 33 (1983)
-
Volume 32 (1982)
-
Volume 31 (1981)
-
Volume 30 (1980)
-
Volume 29 (1979)
-
Volume 28 (1978)
-
Volume 27 (1977)
-
Volume 26 (1976)
-
Volume 25 (1975)
-
Volume 24 (1974)
-
Volume 23 (1973)
-
Volume 22 (1972)
-
Volume 21 (1971)
-
Volume 20 (1970)
-
Volume 19 (1969)
-
Volume 18 (1968)
-
Volume 17 (1967)
-
Volume 16 (1966)
-
Volume 15 (1965)
-
Volume 14 (1964)
-
Volume 13 (1963)
-
Volume 12 (1962)
-
Volume 11 (1961)
-
Volume 10 (1960)
-
Volume 9 (1959)
-
Volume 8 (1958)
-
Volume 7 (1957)
-
Volume 6 (1956)
-
Volume 5 (1955)
-
Volume 4 (1954)
-
Volume 3 (1953)
-
Volume 2 (1952)
-
Volume 1 (1951)
Most Read This Month


