Coronaviruses
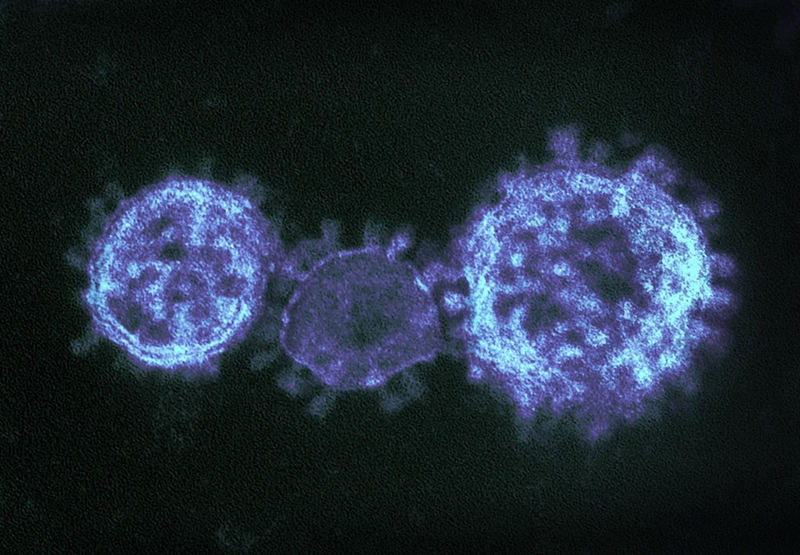
Coronaviruses are a large family of viruses that can infect a range of hosts. They are known to cause diseases including the common cold, Severe Acute Respiratory Syndrome (SARS) and Middle East Respiratory Syndrome (MERS) in humans.
In January 2020, China saw an outbreak of a new coronavirus strain now named SARS-CoV-2. Although the animal reservoir for the SARS and MERS viruses are known, this has yet to have been confirmed for SARS-CoV-2. All three strains are transmissible between humans.
To allow the widest possible distribution of relevant research, the Microbiology Society has brought together articles from across our portfolio and made this content freely available.
Image credit: "MERS-CoV" by NIAID is licensed under CC BY 2.0, this image has been modified.
Collection Contents
151 - 200 of 298 results
-
-
Type I feline coronavirus spike glycoprotein fails to recognize aminopeptidase N as a functional receptor on feline cell lines
More LessThere are two types of feline coronaviruses that can be distinguished by serology and sequence analysis. Type I viruses, which are prevalent in the field but are difficult to isolate and propagate in cell culture, and type II viruses, which are less prevalent but replicate well in cell culture. An important determinant of coronavirus infection, in vivo and in cell culture, is the interaction of the virus surface glycoprotein with a cellular receptor. It is generally accepted that feline aminopeptidase N can act as a receptor for the attachment and entry of type II strains, and it has been proposed that the same molecule acts as a receptor for type I viruses. However, the experimental data are inconclusive. The aim of the studies reported here was to provide evidence for or against the involvement of feline aminopeptidase N as a receptor for type I feline coronaviruses. Our approach was to produce retroviral pseudotypes that bear the type I or type II feline coronavirus surface glycoprotein and to screen a range of feline cell lines for the expression of a functional receptor for attachment and entry. Our results show that type I feline coronavirus surface glycoprotein fails to recognize feline aminopeptidase N as a functional receptor on three continuous feline cell lines. This suggests that feline aminopeptidase N is not a receptor for type I feline coronaviruses. Our results also indicate that it should be possible to use retroviral pseudotypes to identify and characterize the cellular receptor for type I feline coronaviruses.
-
-
-
Molecular analysis of the S glycoprotein gene of bovine coronaviruses isolated in Japan from 1999 to 2006
More LessIn total, 55 isolates of Bovine coronavirus (BCoV) were collected from cases of enteric and respiratory disease occurring between 1999 and 2006 in Japan. Phylogenetic analysis of the polymorphic region of the S glycoprotein gene of these isolates, together with those of other known strains, classified the BCoV strains and isolates into four clusters. Recent field isolates display distinctive genetic divergence from the prototype enteric BCoV strains – Mebus, Quebec, Kakegawa, F15 and LY138 – and have diverged in three different aspects over 8 years. These data suggested that the genetic divergence in the polymorphic region of the S glycoprotein has progressed considerably; thus, molecular analysis of this region should be useful in investigating the molecular epidemiology of BCoV. In addition, based on the differences in amino acids among the isolates, our study did not reveal the presence of certain genetic markers of pathogenicity and clinical symptoms in this polymorphic region.
-
-
-
Full-length genome sequences of two SARS-like coronaviruses in horseshoe bats and genetic variation analysis
More LessBats were recently identified as natural reservoirs of SARS-like coronavirus (SL-CoV) or SARS coronavirus-like virus. These viruses, together with SARS coronaviruses (SARS-CoV) isolated from human and palm civet, form a distinctive cluster within the group 2 coronaviruses of the genus Coronavirus, tentatively named group 2b (G2b). In this study, complete genome sequences of two additional group 2b coronaviruses (G2b-CoVs) were determined from horseshoe bat Rhinolophus ferrumequinum (G2b-CoV Rf1) and Rhinolophus macrotis (G2b-CoV Rm1). The bat G2b-CoV isolates have an identical genome organization and share an overall genome sequence identity of 88–92 % among themselves and between them and the human/civet isolates. The most variable regions are located in the genes encoding nsp3, ORF3a, spike protein and ORF8 when bat and human/civet G2b-CoV isolates are compared. Genetic analysis demonstrated that a diverse G2b-CoV population exists in the bat habitat and has evolved from a common ancestor of SARS-CoV.
-
-
-
Inter- and intra-variant genetic heterogeneity of human coronavirus OC43 strains in France
More LessHuman coronavirus OC43 (HCoV-OC43) causes acute, self-limited respiratory infections. A close relationship between bovine coronaviruses (BCoVs) and HCoV-OC43 has recently been demonstrated. This study includes seven clinical, non-cell culture-adapted, contemporary HCoV-OC43 strains detected in France in 2003. By using RT-PCR and clonal sequencing of the S1 gene of HCoV-OC43, the inter-variant heterogeneity of the HCoV-OC43 circulating strains was studied and the intra-variant diversity was assessed by investigation of a quasispecies cloud. This paper brings to the forefront a high genetic diversity of circulating HCoV-OC43 variants. Genetically different groups are defined among the variants described in this study. One of these variants holds characteristics of an outlier and presents a deletion of 12 nt, also found in BCoV strains. Moreover, the presence of HCoV-OC43 as a quasispecies cloud in vivo during an acute respiratory-tract illness was discovered. It has also been revealed that quasispecies-cloud sizes are similar for the two viral populations tested.
-
-
-
Coronaviruses in bent-winged bats (Miniopterus spp.)
More LessA novel group 1 coronavirus was previously identified in bent-winged bats (Miniopterus spp.). Here, results are described from our ongoing surveillance of these bats for coronaviruses. These findings show that group 1 coronaviruses are endemic in these bat populations in Hong Kong. Genetic analysis of these viruses indicates that there are at least four different, but closely related, group 1 coronaviruses (bat-CoV 1A, 1B, HKU7 and HKU8) circulating in bent-winged bats. Phylogenetic analysis revealed that these group 1 bat coronaviruses have descended from a common ancestor and that these viruses have been established in these bats for a long period of time. These data provide a better understanding of the emergence and evolution of coronaviruses. Bat-CoV 1A and 1B were detected in apparently healthy Miniopterus magnater and Miniopterus pusillus, respectively, on repeated sampling occasions at a single habitat, suggesting that these viruses have established a persistent infection in these populations.
-
-
-
Interaction of severe acute respiratory syndrome-associated coronavirus with dendritic cells
More LessSevere acute respiratory syndrome (SARS) of humans is caused by a novel coronavirus of zoonotic origin termed SARS-associated coronavirus (SARS-CoV). The virus induces severe injury of lung tissue, as well as lymphopenia and destruction of the architecture of lymphatic tissue by as-yet-unknown mechanisms. In this study, the interaction of SARS-CoV with dendritic cells (DCs), the key regulators of immune responses, was analysed. Monocyte-derived DCs were infected with SARS-CoV and analysed for viability, surface-marker expression and alpha interferon (IFN-α) induction. SARS-CoV infection was monitored by quantitative RT-PCR, immunofluorescence analysis and recovery experiments. SARS-CoV infected both immature and mature DCs, although replication efficiency was low. Immature DCs were activated by SARS-CoV infection and by UV-inactivated SARS-CoV. Infected DCs were still viable on day 6 post-infection, but major histocompatibility complex class I upregulation was missing, indicating that DC function was impaired. Additionally, SARS-CoV infection induced a delayed activation of IFN-α expression. Therefore, it is concluded that SARS-CoV has the ability to circumvent both the innate and the adaptive immune systems.
-
-
-
Analysis of ACE2 in polarized epithelial cells: surface expression and function as receptor for severe acute respiratory syndrome-associated coronavirus
More LessThe primary target of severe acute respiratory syndrome-associated coronavirus (SARS-CoV) is epithelial cells in the respiratory and intestinal tract. The cellular receptor for SARS-CoV, angiotensin-converting enzyme 2 (ACE2), has been shown to be localized on the apical plasma membrane of polarized respiratory epithelial cells and to mediate infection from the apical side of these cells. Here, these results were confirmed and extended by including a colon carcinoma cell line (Caco-2), a lung carcinoma cell line (Calu-3) and Vero E6 cells in our analysis. All three cell types expressed human ACE2 on the apical membrane domain and were infected via this route, as determined with vesicular stomatitis virus pseudotypes containing the S protein of SARS-CoV. In a histological analysis of the respiratory tract, ACE2 was detected in the trachea, main bronchus and alveoli, and occasionally also in the small bronchi. These data will help us to understand the pathogenesis of SARS-CoV infection.
-
-
-
Analysis of human coronavirus 229E spike and nucleoprotein genes demonstrates genetic drift between chronologically distinct strains
More LessHistorically, coronaviruses have been recognized as a cause of minor respiratory infections in humans. However, the recent identification of three novel human coronaviruses, one causing severe acute respiratory syndrome (SARS), has prompted further examination of these viruses. Previous studies of geographically and chronologically distinct Human coronavirus 229E (HCoV-229E) isolates have found only limited variation within S gene nucleotide sequences. In contrast, analysis of the S genes of contemporary Human coronavirus OC43 variants identified in Belgium revealed two distinct viruses circulating during 2003 and 2004. Here, the S and N gene sequences of 25 HCoV-229E variants identified in Victoria, Australia, between 1979 and 2004 in patients with symptomatic infections were determined. Phylogenetic analysis showed clustering of the isolates into four groups, with evidence of increasing divergence with time. Evidence of positive selection in the S gene was also established.
-
-
-
Comparative evaluation of two severe acute respiratory syndrome (SARS) vaccine candidates in mice challenged with SARS coronavirus
More LessRaymond H. See, Alexander N. Zakhartchouk, Martin Petric, David J. Lawrence, Catherine P. Y. Mok, Robert J. Hogan, Thomas Rowe, Lois A. Zitzow, Karuna P. Karunakaran, Mary M. Hitt, Frank L. Graham, Ludvik Prevec, James B. Mahony, Chetna Sharon, Thierry C. Auperin, James M. Rini, Aubrey J. Tingle, David W. Scheifele, Danuta M. Skowronski, David M. Patrick, Thomas G. Voss, Lorne A. Babiuk, Jack Gauldie, Rachel L. Roper, Robert C. Brunham and B. Brett FinlayTwo different severe acute respiratory syndrome (SARS) vaccine strategies were evaluated for their ability to protect against live SARS coronavirus (CoV) challenge in a murine model of infection. A whole killed (inactivated by β-propiolactone) SARS-CoV vaccine and a combination of two adenovirus-based vectors, one expressing the nucleocapsid (N) and the other expressing the spike (S) protein (collectively designated Ad S/N), were evaluated for the induction of serum neutralizing antibodies and cellular immune responses and their ability to protect against pulmonary SARS-CoV replication. The whole killed virus (WKV) vaccine given subcutaneously to 129S6/SvEv mice was more effective than the Ad S/N vaccine administered either intranasally or intramuscularly in inhibiting SARS-CoV replication in the murine respiratory tract. This protective ability of the WKV vaccine correlated with the induction of high serum neutralizing-antibody titres, but not with cellular immune responses as measured by gamma interferon secretion by mouse splenocytes. Titres of serum neutralizing antibodies induced by the Ad S/N vaccine administered intranasally or intramuscularly were significantly lower than those induced by the WKV vaccine. However, Ad S/N administered intranasally, but not intramuscularly, significantly limited SARS-CoV replication in the lungs. Among the vaccine groups, SARS-CoV-specific IgA was found only in the sera of mice immunized intranasally with Ad S/N, suggesting that mucosal immunity may play a role in protection for the intranasal Ad S/N delivery system. Finally, the sera of vaccinated mice contained antibodies to S, further suggesting a role for this protein in conferring protective immunity against SARS-CoV infection.
-
-
-
Amino terminus of the SARS coronavirus protein 3a elicits strong, potentially protective humoral responses in infected patients
More LessThe 3a protein of severe acute respiratory syndrome (SARS)-associated coronavirus is expressed and transported to the plasma membrane in tissue cells of infected patients. Its short N-terminal ectodomain was found to elicit strong humoral responses in half of the patients who had recovered from SARS. The ectodomain-specific antibodies from the convalescent-phase plasma readily recognized and induced destruction of 3a-expressing cells in the presence of the human complement system, demonstrating their potential ability to provide immune protection by recognizing and eliminating SARS coronavirus-infected cells that express the target protein. In addition, when coupled to a carrier protein, the ectodomain peptide elicited 3a-specific antibodies in mice and rabbit at high titres. These results showed that the N terminus of the 3a protein is highly immunogenic and elicits potentially protective humoral responses in infected patients. Therefore, the short extracellular domain may be a valuable immunogen in the development of a vaccine for infectious SARS.
-
-
-
Subcellular localization of the severe acute respiratory syndrome coronavirus nucleocapsid protein
More LessThe coronavirus nucleocapsid (N) protein is a viral RNA-binding protein with multiple functions in terms of virus replication and modulating cell signalling pathways. N protein is composed of three distinct regions containing RNA-binding motif(s), and appropriate signals for modulating cell signalling. The subcellular localization of severe acute respiratory syndrome coronavirus (SARS-CoV) N protein was studied. In infected cells, SARS-CoV N protein localized exclusively to the cytoplasm. In contrast to the avian coronavirus N protein, overexpressed SARS-CoV N protein remained principally localized to the cytoplasm, with very few cells exhibiting nucleolar localization. Bioinformatic analysis and deletion mutagenesis coupled to confocal microscopy and live-cell imaging, revealed that SARS-CoV N protein regions I and III contained nuclear localization signals and region II contained a nucleolar retention signal. However, cytoplasmic localization was directed by region III and was the dominant localization signal in the protein.
-
-
-
Genomic RNA sequence of Feline coronavirus strain FIPV WSU-79/1146
More LessA consensus sequence of the Feline coronavirus (FCoV) (strain FIPV WSU-79/1146) genome was determined from overlapping cDNA fragments produced by RT-PCR amplification of viral RNA. The genome was found to be 29 125 nt in length, excluding the poly(A) tail. Analysis of the sequence identified conserved open reading frames and revealed an overall genome organization similar to that of other coronaviruses. The genomic RNA was analysed for putative cis-acting elements and the pattern of subgenomic mRNA synthesis was analysed by Northern blotting. Comparative sequence analysis of the predicted FCoV proteins identified 16 replicase proteins (nsp1–nsp16) and four structural proteins (spike, membrane, envelope and nucleocapsid). Two mRNAs encoding putative accessory proteins were also detected. Phylogenetic analyses confirmed that FIPV WSU-79/1146 belongs to the coronavirus subgroup G1-1. These results confirm and extend previous findings from partial sequence analysis of FCoV genomes.
-
-
-
Vesicular stomatitis virus pseudotyped with severe acute respiratory syndrome coronavirus spike protein
More LessSevere acute respiratory syndrome coronavirus (SARS-CoV) contains a single spike (S) protein, which binds to its receptor, angiotensin-converting enzyme 2 (ACE2), induces membrane fusion and serves as a neutralizing antigen. A SARS-CoV-S protein-bearing vesicular stomatitis virus (VSV) pseudotype using the VSVΔG* system was generated. Partial deletion of the SARS-CoV-S protein cytoplasmic domain allowed efficient incorporation into VSV particles and led to the generation of a pseudotype (VSV-SARS-St19) at high titre. Green fluorescent protein expression was demonstrated as early as 7 h after infection of Vero E6 cells with VSV-SARS-St19. VSV-SARS-St19 was neutralized by anti-SARS-CoV antibody and soluble ACE2, and its infection was blocked by treatment of Vero E6 cells with anti-ACE2 antibody. These results indicated that VSV-SARS-St19 infection is mediated by SARS-CoV-S protein in an ACE2-dependent manner. VSV-SARS-St19 will be useful for analysing the function of SARS-CoV-S protein and for developing rapid methods of detecting neutralizing antibodies specific for SARS-CoV infection.
-
-
-
The 3a protein of severe acute respiratory syndrome-associated coronavirus induces apoptosis in Vero E6 cells
More LessAn outbreak of severe acute respiratory syndrome (SARS) occurred in China and the first case emerged in mid-November 2002. The aetiological agent of this disease was found to be a previously unknown coronavirus, SARS-associated coronavirus (SARS-CoV). The detailed pathology of SARS-CoV infection and the host response to the viral infection are still not known. The 3a gene encodes a non-structural viral protein, which is predicted to be a transmembrane protein. In this study, it was shown that the 3a protein was expressed in the lungs and intestinal tissues of SARS patients and that the protein localized to the endoplasmic reticulum in 3a-transfected monkey kidney Vero E6 cells. In vitro experiments of chromatin condensation and DNA fragmentation suggested that the 3a protein may trigger apoptosis. These data showed that overexpression of a single SARS-CoV protein can induce apoptosis in vitro.
-
-
-
Molecular identification and characterization of novel coronaviruses infecting graylag geese (Anser anser), feral pigeons (Columbia livia) and mallards (Anas platyrhynchos)
More LessIn light of the finding of a previously unknown coronavirus as the aetiology of the severe acute respiratory syndrome (SARS), it is probable that other coronaviruses, than those recognized to date, are circulating in animal populations. Here, the results of a screening for coronavirus are presented, using a universal coronavirus RT-PCR, of the bird species graylag goose (Anser anser), feral pigeon (Columbia livia) and mallard (Anas platyrhynchos). Coronaviruses were found in cloacal swab samples from all the three bird species. In the graylag goose, 40 of 163 sampled birds were coronavirus positive, whereas two of 100 sampled pigeons and one of five sampled mallards tested positive. The infected graylag geese showed lower body weights compared with virus-negative birds, suggesting clinical significance of the infection. Phylogenetic analyses performed on the replicase gene and nucleocapsid protein sequences, indicated that the novel coronaviruses described in the present study all branch off from group III coronaviruses. All the novel avian coronaviruses harboured the conserved s2m RNA structure in their 3′ untranslated region, like other previously described group III coronaviruses, and like the SARS coronavirus. Sequencing of the complete nucleocapsid gene and downstream regions of goose and pigeon coronaviruses, evidenced the presence of two additional open reading frames for the goose coronavirus with no sequence similarity to known proteins, but with predicted transmembrane domains for one of the encoded proteins, and one additional open reading frame for the pigeon coronavirus, with a predicted transmembrane domain, downstream of the nucleocapsid gene.
-
-
-
A single immunization with a rhabdovirus-based vector expressing severe acute respiratory syndrome coronavirus (SARS-CoV) S protein results in the production of high levels of SARS-CoV-neutralizing antibodies
More LessForeign viral proteins expressed by rabies virus (RV) have been shown to induce potent humoral and cellular immune responses in immunized animals. In addition, highly attenuated and, therefore, very safe RV-based vectors have been constructed. Here, an RV-based vaccine vehicle was utilized as a novel vaccine against severe acute respiratory syndrome coronavirus (SARS-CoV). For this approach, the SARS-CoV nucleocapsid protein (N) or envelope spike protein (S) genes were cloned between the RV glycoprotein G and polymerase L genes. Recombinant vectors expressing SARS-CoV N or S protein were recovered and their immunogenicity was studied in mice. A single inoculation with the RV-based vaccine expressing SARS-CoV S protein induced a strong SARS-CoV-neutralizing antibody response. The ability of the RV-SARS-CoV S vector to confer immunity after a single inoculation makes this live vaccine a promising candidate for eradication of SARS-CoV in animal reservoirs, thereby reducing the risk of transmitting the infection to humans.
-
-
-
Differential maturation and subcellular localization of severe acute respiratory syndrome coronavirus surface proteins S, M and E
More LessPost-translational modifications and correct subcellular localization of viral structural proteins are prerequisites for assembly and budding of enveloped viruses. Coronaviruses, like the severe acute respiratory syndrome-associated virus (SARS-CoV), bud from the endoplasmic reticulum-Golgi intermediate compartment. In this study, the subcellular distribution and maturation of SARS-CoV surface proteins S, M and E were analysed by using C-terminally tagged proteins. As early as 30 min post-entry into the endoplasmic reticulum, high-mannosylated S assembles into trimers prior to acquisition of complex N-glycans in the Golgi. Like S, M acquires high-mannose N-glycans that are subsequently modified into complex N-glycans in the Golgi. The N-glycosylation profile and the absence of O-glycosylation on M protein relate SARS-CoV to the previously described group 1 and 3 coronaviruses. Immunofluorescence analysis shows that S is detected in several compartments along the secretory pathway from the endoplasmic reticulum to the plasma membrane while M predominantly localizes in the Golgi, where it accumulates, and in trafficking vesicles. The E protein is not glycosylated. Pulse-chase labelling and confocal microscopy in the presence of protein translation inhibitor cycloheximide revealed that the E protein has a short half-life of 30 min. E protein is found in bright perinuclear patches colocalizing with endoplasmic reticulum markers. In conclusion, SARS-CoV surface proteins S, M and E show differential subcellular localizations when expressed alone suggesting that additional cellular or viral factors might be required for coordinated trafficking to the virus assembly site in the endoplasmic reticulum-Golgi intermediate compartment.
-
-
-
Isolation of avian infectious bronchitis coronavirus from domestic peafowl (Pavo cristatus) and teal (Anas)
More LessCoronavirus-like viruses, designated peafowl/China/LKQ3/2003 (pf/CH/LKQ3/03) and teal/China/LDT3/2003 (tl/CH/LDT3/03), were isolated from a peafowl and a teal during virological surveillance in Guangdong province, China. Partial genomic sequence analysis showed that these isolates had the S–3–M–5–N gene order that is typical of avian coronaviruses. The spike, membrane and nucleocapsid protein genes of pf/CH/LKQ3/03 had >99 % identity to those of the avian infectious bronchitis coronavirus H120 vaccine strain (Massachusetts serotype) and other Massachusetts serotype isolates. Furthermore, when pf/CH/LKQ3/03 was inoculated experimentally into chickens (specific-pathogen-free), no disease signs were apparent. tl/CH/LDT3/03 had a spike protein gene with 95 % identity to that of a Chinese infectious bronchitis virus (IBV) isolate, although more extensive sequencing revealed the possibility that this strain may have undergone recombination. When inoculated into chickens, tl/CH/LDT3/03 resulted in the death of birds from nephritis. Taken together, this information suggests that pf/CH/LKQ3/03 might be a revertant, attenuated vaccine IBV strain, whereas tl/CH/LDT3/03 is a nephropathogenic field IBV strain, generated through recombination. The replication and non-pathogenic nature of IBV in domestic peafowl and teal under field conditions raises questions as to the role of these hosts as carriers of IBV and the potential that they may have to transmit virus to susceptible chicken populations.
-
-
-
Severe acute respiratory syndrome coronavirus nucleocapsid protein expressed by an adenovirus vector is phosphorylated and immunogenic in mice
More LessSevere acute respiratory syndrome coronavirus (SARS-CoV) has been identified as the aetiological agent of SARS. Thus, vaccination against SARS-CoV may represent an effective approach towards controlling SARS. The nucleocapsid (N) protein is thought to play a role in induction of cell-mediated immunity to SARS-CoV and thus it is important to characterize this protein. In the present study, an E1/partially E3-deleted, replication-defective human adenovirus 5 (Ad5) vector (Ad5-N-V) expressing the SARS-CoV N protein was constructed. The N protein, expressed in vitro by Ad5-N-V, was of the expected molecular mass of 50 kDa and was phosphorylated. Vaccination of C57BL/6 mice with Ad5-N-V generated potent SARS-CoV-specific humoral and T cell-mediated immune responses. These results show that Ad5-N-V may potentially be used as a SARS-CoV vaccine.
-
-
-
Rapid identification of coronavirus replicase inhibitors using a selectable replicon RNA
More LessA previously unknown coronavirus (CoV) is the aetiological agent causing severe acute respiratory syndrome (SARS), for which an effective antiviral treatment is urgently needed. To enable the rapid and biosafe identification of coronavirus replicase inhibitors, we have generated a non-cytopathic, selectable replicon RNA (based on human CoV 229E) that can be stably maintained in eukaryotic cells. Most importantly, the replicon RNA mediates reporter gene expression as a marker for coronavirus replication. We have used a replicon RNA-containing cell line to test the inhibitory effect of several compounds that are currently being assessed for SARS treatment. Amongst those, interferon-α displayed the strongest inhibitory activity. Our results demonstrate that coronavirus replicon cell lines provide a versatile and safe assay for the identification of coronavirus replicase inhibitors. Once this technology is adapted to SARS-CoV replicon RNAs, it will allow high throughput screening for SARS-CoV replicase inhibitors without the need to grow infectious SARS-CoV.
-
-
-
Antibody response and viraemia during the course of severe acute respiratory syndrome (SARS)-associated coronavirus infection
More LessTo understand the time-course of viraemia and antibody responses to severe acute respiratory syndrome-associated coronavirus (SARS-CoV), RT-PCR and ELISA were used to assay 376 blood samples from 135 SARS patients at various stages of the illness, including samples from patients who were in their early convalescent phase. The results showed that IgM antibodies decreased and became undetectable 11 weeks into the recovery phase. IgG antibodies, however, remained detectable for a period beyond 11 weeks and were found in 100 % of patients in the early convalescent phase. SARS-CoV viraemia mainly appeared 1 week after the onset of illness and then decreased over a period of 1 month, becoming undetectable in the blood samples of the convalescent patients. At the peak of viraemia, viral RNA was detectable in 75 % of blood samples from patients who were clinically diagnosed with SARS 1 or 2 weeks before the test.
-
-
-
Proliferative growth of SARS coronavirus in Vero E6 cells
More LessAn isolate of SARS coronavirus (strain 2003VA2774) was obtained from a patient and used to infect Vero E6 cells. The replication cycle of the virus was followed from 1 to 30 h post-infection (p.i.). It was surprising to observe the swift growth of this human virus in Vero cells. Within the first hour of infection, the most obvious ultrastructural change was the proliferation of the Golgi complexes and related vesicles accompanied by swelling of some of the trans-Golgi sacs. Extracellular virus particles were present by 5 h p.i. in about 5 % of the cells and this increased dramatically to about 30 % of the cell population within an hour (6 h p.i.). Swollen Golgi sacs contained virus nucleocapsids at different stages of maturation. These virus precursors were also in large vacuoles and in close association with membrane whorls. The membrane whorls could be the replication complexes, since they appeared rather early in the replication cycle. As infection progressed from 12 to 21 h p.i., the cytoplasm of the infected cells was filled with numerous large, smooth-membraned vacuoles containing a mixture of mature virus and spherical cores. Several of these vacuoles were close to the cell periphery, ready to export out the mature progeny virus particles via exocytosis. By 24 to 30 h p.i., crystalline arrays of the extracellular virus particles were seen commonly at the cell surface.
-
-
-
Persistence and transmission of natural type I feline coronavirus infection
More LessTo examine the mode of natural transmission and persistence of feline coronavirus (FCoV), FCoV strains shed by domestic cats were investigated over periods of up to 7 years. An RT-PCR that amplified part of the 3′ end of the viral spike (S) gene was devised to distinguish FCoV types I and II. All but 1 of 28 strains of FCoV from 43 cats were type I. Nucleotide identities of the amplified 320 bp product from 49 type I FCoVs ranged from 79 to 100 %. The consensus partial S sequence of isolates recovered from persistently infected cats at time intervals spanning years was generally conserved. While most cats were infected with a single strain, a few may have been infected by more than one strain. Cats that were transiently infected and ceased shedding could be re-infected with either the same, or a different, strain. In most cases, whether a cat became persistently or transiently infected was independent of the virus strain. However, one strain was unusual in that it infected the majority of cats in the household simultaneously and was still being shed 18 months later. Factors that influence whether FCoV establishes lifelong infection in some cats and not others are determined mainly by the host response to infection.
-
-
-
Mechanisms and enzymes involved in SARS coronavirus genome expression
More LessA novel coronavirus is the causative agent of the current epidemic of severe acute respiratory syndrome (SARS). Coronaviruses are exceptionally large RNA viruses and employ complex regulatory mechanisms to express their genomes. Here, we determined the sequence of SARS coronavirus (SARS-CoV), isolate Frankfurt 1, and characterized key RNA elements and protein functions involved in viral genome expression. Important regulatory mechanisms, such as the (discontinuous) synthesis of eight subgenomic mRNAs, ribosomal frameshifting and post-translational proteolytic processing, were addressed. Activities of three SARS coronavirus enzymes, the helicase and two cysteine proteinases, which are known to be critically involved in replication, transcription and/or post-translational polyprotein processing, were characterized. The availability of recombinant forms of key replicative enzymes of SARS coronavirus should pave the way for high-throughput screening approaches to identify candidate inhibitors in compound libraries.
-
-
-
Sequence of the 3′-terminal end (8·1 kb) of the genome of porcine haemagglutinating encephalomyelitis virus: comparison with other haemagglutinating coronaviruses
More LessA cytopathogenic coronavirus, serologically identified as porcine haemagglutinating encephalomyelitis virus (HEV), has recently been associated with acute outbreaks of wasting and encephalitis in nursing piglets from pig farms in southern Québec and Ontario, Canada. The 3′-terminal end of the genome of the prototype HEV-67N strain and that of the recent Québec IAF-404 field isolate, both propagated in HRT-18 cells, were sequenced. Overall, sequencing data indicated that HEV has remained antigenically and genetically stable since its first isolation in North America in 1962. Compared with the prototype strain of bovine enteropathogenic coronavirus (BCoV), HEV, as well as the human respiratory coronavirus (HCoV-OC43) showed a major deletion in their ORF4 gene. Deduced amino acid sequences for both HEV strains revealed 89/88, 80, 93/92 and 95/94% identities with the structural proteins HE, S, M and N of BCoV and HCoV-OC43, respectively. Major variations were observed in the S1 portion of the S gene of both HEV strains, with only 73/71% amino acid identities compared with those of the two other haemagglutinating coronaviruses.
-
-
-
Conservation of substrate specificities among coronavirus main proteases
More LessThe key enzyme in coronavirus replicase polyprotein processing is the coronavirus main protease, 3CLpro. The substrate specificities of five coronavirus main proteases, including the prototypic enzymes from the coronavirus groups I, II and III, were characterized. Recombinant main proteases of human coronavirus (HCoV), transmissible gastroenteritis virus (TGEV), feline infectious peritonitis virus, avian infectious bronchitis virus and mouse hepatitis virus (MHV) were tested in peptide-based trans-cleavage assays. The determination of relative rate constants for a set of corresponding HCoV, TGEV and MHV 3CLpro cleavage sites revealed a conserved ranking of these sites. Furthermore, a synthetic peptide representing the N-terminal HCoV 3CLpro cleavage site was shown to be effectively hydrolysed by noncognate main proteases. The data show that the differential cleavage kinetics of sites within pp1a/pp1ab are a conserved feature of coronavirus main proteases and lead us to predict similar processing kinetics for the replicase polyproteins of all coronaviruses.
-
-
-
Mutational analysis of the active centre of coronavirus 3C-like proteases
More LessFormation of the coronavirus replication–transcription complex involves the synthesis of large polyprotein precursors that are extensively processed by virus-encoded cysteine proteases. In this study, the coding sequence of the feline infectious peritonitis virus (FIPV) main protease, 3CLpro, was determined. Comparative sequence analyses revealed that FIPV 3CLpro and other coronavirus main proteases are related most closely to the 3C-like proteases of potyviruses. The predicted active centre of the coronavirus enzymes has accepted unique replacements that were probed by extensive mutational analysis. The wild-type FIPV 3CLpro domain and 25 mutants were expressed in Escherichia coli and tested for proteolytic activity in a peptide-based assay. The data strongly suggest that, first, the FIPV 3CLpro catalytic system employs His41 and Cys144 as the principal catalytic residues. Second, the amino acids Tyr160 and His162, which are part of the conserved sequence signature Tyr160–Met161–His162 and are believed to be involved in substrate recognition, were found to be indispensable for proteolytic activity. Third, replacements of Gly83 and Asn64, which were candidates to occupy the position spatially equivalent to that of the catalytic Asp residue of chymotrypsin-like proteases, resulted in proteolytically active proteins. Surprisingly, some of the Asn64 mutants even exhibited strongly increased activities. Similar results were obtained for human coronavirus (HCoV) 3CLpro mutants in which the equivalent Asn residue (HCoV 3CLpro Asn64) was substituted. These data lead us to conclude that both the catalytic systems and substrate-binding pockets of coronavirus main proteases differ from those of other RNA virus 3C and 3C-like proteases.
-
-
-
The sialate-4-O-acetylesterases of coronaviruses related to mouse hepatitis virus: a proposal to reorganize group 2 Coronaviridae
More LessGroup 2 coronaviruses are characterized within the order Nidovirales by a unique genome organization. A characteristic feature of group 2 coronaviruses is the presence of a gene encoding the haemagglutinin–esterase (HE) protein, which is absent in coronaviruses of groups 1 and 3. At least three coronavirus strains within group 2 expressed a structural protein with sialate-4-O-acetylesterase activity, distinguishing them from other members of group 2, which encode an enzyme specific for 5-N-acetyl-9-O-acetylneuraminic acid. The esterases of mouse hepatitis virus (MHV) strains S and JHM and puffinosis virus (PV) specifically hydrolysed 5-N-acetyl-4-O-acetylneuraminic acid (Neu4,5Ac2) as well as the synthetic substrates p-nitrophenyl acetate, 4-methylumbelliferyl acetate and fluorescein diacetate. The K m values of the MHV-like esterases for the latter substrates were two- to tenfold lower than those of the sialate-9-O-acetylesterases of influenza C viruses. Another unspecific esterase substrate, α-naphthyl acetate, was used for the in situ detection of the dimeric HE proteins in SDS–polyacrylamide gels. MHV-S, MHV-JHM and PV bound to horse serum glycoproteins containing Neu4,5Ac2. De-O-acetylation of the glycoproteins by alkaline treatment or incubation with the viral esterases resulted in a complete loss of recognition, indicating a specific interaction of MHV-like coronaviruses with Neu4,5Ac2. Combined with evidence for distinct phylogenetic lineages of group 2 coronaviruses, subdivision into subgroups 2a (MHV-like viruses) and 2b (bovine coronavirus-like viruses) is suggested.
-
-
-
Comparison of genomic and predicted amino acid sequences of respiratory and enteric bovine coronaviruses isolated from the same animal with fatal shipping pneumonia
More LessThe complete genome sequences are reported here of two field isolates of bovine coronavirus (BCoV), which were isolated from respiratory and intestinal samples of the same animal experiencing fatal pneumonia during a bovine shipping fever epizootic. Both genomes contained 31028 nucleotides and included 13 open reading frames (ORFs) flanked by 5′- and 3′-untranslated regions (UTRs). ORF1a and ORF1b encode replicative polyproteins pp1a and pp1ab, respectively, that contain all of the putative functional domains documented previously for the closest relative, mouse hepatitis virus. The genomes of the BCoV isolates differed in 107 positions, scattered throughout the genome except the 5′-UTR. Differences in 25 positions were non-synonymous and were located in all proteins except pp1b. Six replicase mutations were identified within or immediately downstream of the predicted largest pp1a-derived protein, p195/p210. Single amino acid changes within p195/p210 as well as within the S glycoprotein might contribute to the different phenotypes of the BCoV isolates.
-
-
-
Infectious RNA transcribed in vitro from a cDNA copy of the human coronavirus genome cloned in vaccinia virus
More LessThe coronavirus genome is a positive-strand RNA of extraordinary size and complexity. It is composed of approximately 30000 nucleotides and it is the largest known autonomously replicating RNA. It is also remarkable in that more than two-thirds of the genome is devoted to encoding proteins involved in the replication and transcription of viral RNA. Here, a reverse-genetic system is described for the generation of recombinant coronaviruses. This system is based upon the in vitro transcription of infectious RNA from a cDNA copy of the human coronavirus 229E genome that has been cloned and propagated in vaccinia virus. This system is expected to provide new insights into the molecular biology and pathogenesis of coronaviruses and to serve as a paradigm for the genetic analysis of large RNA virus genomes. It also provides a starting point for the development of a new class of eukaryotic, multi-gene RNA vectors that are able to express several proteins simultaneously.
-
-
-
Functional analysis of an epitope in the S2 subunit of the murine coronavirus spike protein: involvement in fusion activity
More LessThe monoclonal antibody (MAb) 5B19.2, which has virus-neutralizing and fusion inhibition activities, binds to an epitope (S2A) consisting of nine hydrophobic amino acids in the S2 subunit of the mouse hepatitis virus (MHV) spike (S) protein. This suggests that the S2A epitope may be involved in binding the virus to the MHV receptor and/or in virus–cell fusion. Co-immunoprecipitation analyses demonstrated that while the binding of virus to the receptor was blocked by anti-S1 MAbs, it was not blocked by the S2A antiserum, indicating that S2A was not involved in receptor-binding. The S proteins prepared in this study with mutations in the S2A epitope were either fusogenic or non-fusogenic and their fusogenicity did not correlate with the hydrophobic feature of the S2A epitope. All of these wt and mutated S proteins were similarly transported onto the cell membrane independent of their fusogenicity capability. These results suggest that S2A may mediate the fusion activity of the MHV S protein during virus entry into cells.
-
-
-
Temperature-sensitive acetylesterase activity of haemagglutinin-esterase specified by respiratory bovine coronaviruses
More LessNumerous respiratory bovine coronaviruses (RBCV) were isolated recently from nasal swab samples and lung tissues of feedlot cattle with acute respiratory tract disease. These newly emerging RBCV isolates exhibited distinct phenotypic features that differentiated them from enteropathogenic bovine coronaviruses (EBCV). The RBCV strains had a receptor-destroying enzyme function mediated by acetylesterase (AE) activity of the haemagglutinin-esterase (HE) glycoprotein. The HE genes of wild-type EBCV strain LY138 and RBCV strains OK-0514 (OK) and LSU-94LSS-051 (LSU) were cloned, sequenced and transiently expressed in COS-7 cells. The enzymic properties of HE proteins in COS-7 cellular extracts and in purified virus preparations were assayed at room temperature, 37°C and 39°C by two different assays. One assay used ρ-nitrophenyl acetate (PNPA) as substrate and detected serine-esterase activity; the second assay monitored AE function with bovine submaxillary mucin (BSM) as substrate. The PNPA tests confirmed that HE proteins of EBCV and RBCV were functionally expressed in transfected COS-7 cells. Time-dependent determination of the AE activity of purified RBCV OK and LSU particles showed lower AE activity at 39°C than at 37°C, whereas the purified EBCV LY particles retained full AE activity at both 37°C and 39°C. Transiently expressed RBCV HE exhibited a marked reduction of AE activity after 40 min of assay time at 37°C. In contrast, the AE activity of the transiently expressed EBCV HE remained stable beyond 40 min. The deduced amino-acid sequences of the HE proteins specified by the RBCV strains OK and LSU contained specific amino-acid changes in comparison with the EBCV LY strain, which may be responsible for the observed enzymic differences. These results are consistent with the hypothesis that RBCV strains have evolved to selectively replicate in respiratory tissues and that HE may play a role in this tissue tropism.
-
-
-
Utilizing fowlpox virus recombinants to generate defective RNAs of the coronavirus infectious bronchitis virus
More LessCoronavirus defective RNAs (D-RNAs) have been used as RNA vectors for the expression of heterologous genes and as vehicles for reverse genetics by modifying coronavirus genomes by targetted recombination. D-RNAs based on the avian coronavirus infectious bronchitis virus (IBV) D-RNA CD-61 have been rescued (replicated and packaged into virions) in a helper virus-dependent manner following electroporation of in vitro-generated T7 transcripts into IBV-infected cells. In order to increase the efficiency of rescue of IBV D-RNAs, cDNAs based on CD-61, under the control of a T7 promoter, were integrated into the fowlpox virus (FPV) genome. The 3′-UTR of the D-RNAs was flanked by a hepatitis delta antigenomic ribozyme and T7 terminator sequence to generate suitable 3′ ends for rescue by helper IBV. Cells were co-infected simultaneously with IBV, the recombinant FPV (rFPV) containing the D-RNA sequence and a second rFPV expressing T7 RNA polymerase for the initial expression of the D-RNA transcript, subsequently rescued by helper IBV. Rescue of rFPV-derived CD-61 occurred earlier and with higher efficiency than demonstrated previously for electroporation of in vitro T7-generated RNA transcripts in avian cells. Rescue of CD-61 was also demonstrated for the first time in mammalian cells. The rescue of rFPV-derived CD-61 by M41 helper IBV resulted in leader switching, in which the Beaudette-type leader sequence on CD-61 was replaced with the M41 leader sequence, confirming that helper IBV virus replicated the rFPV-derived D-RNA. An rFPV-derived D-RNA containing the luciferase gene under the control of an IBV transcription-associated sequence was also rescued and expressed luciferase on serial passage.
-
-
-
A potential role for tumour necrosis factor-α in synergy between porcine respiratory coronavirus and bacterial lipopolysaccharide in the induction of respiratory disease in pigs
More LessThis study examined whether exposure of pigs to both porcine respiratory coronavirus (PRCV) and bacterial lipopolysaccharide (LPS) can potentiate respiratory disease and lung secretion of tumour necrosis factor-α (TNF-α) and interleukin-1 (IL-1). Caesarian-derived colostrum-deprived pigs were inoculated intratracheally with PRCV, with LPS from Escherichia coli O111:B4 (20 μg/kg), or with a combination of the two, and killed at set times after inoculation. Clinical signs, virus replication and (histo)pathological changes in the lungs, percentage of neutrophils and bioactive TNF-α and IL-1 in broncho-alveolar lavage (BAL) fluids were examined. The effects of separate virus or LPS inoculations were subclinical and failed to induce high and sustained cytokine levels. In a preliminary study, pigs were inoculated with PRCV and then with LPS 24 h later and killed sequentially. Severe respiratory disease and significantly enhanced TNF-α titres (208–3601 U/ml versus 40–89 U/ml after LPS only) were seen during the first 12 h after LPS inoculation. IL-1 levels (106–1631 U/ml versus 28–654 U/ml after LPS only) were also increased, but persisted for longer after clinical recovery than TNF-α. In a second study, pigs were inoculated with PRCV and subsequently with LPS at various time intervals ranging from 0 to 24 h, and killed 5 h after inoculation with LPS. A time interval of at least 12 h between inoculations was necessary for prominent respiratory signs to develop. Production of TNF-α, but not IL-1, was also dependent on the time interval between inoculations and was tightly correlated with disease. Lung neutrophil infiltration and pathological changes were comparable after combined PRCV-LPS and single LPS inoculations, and were not associated with disease. These data show that exposure to high endotoxin concentrations in swine buildings can precipitate respiratory disease in PRCV-infected pigs, and that TNF-α is probably an important mediator of these effects. This is the first in-vivo demonstration of synergy between respiratory viruses and LPS.
-
-
-
Expression of reporter genes from the defective RNA CD-61 of the coronavirus infectious bronchitis virus
More LessThe defective RNA (D-RNA) CD-61, derived from the Beaudette strain of the avian coronavirus infectious bronchitis virus (IBV), was used as an RNA vector for the expression of two reporter genes, luciferase and chloramphenicol acetyltransferase (CAT). D-RNAs expressing the CAT gene were demonstrated to be capable of producing CAT protein in a helper-dependent expression system to about 1·6 μg per 106 cells. The reporter genes were expressed from two different sites within the CD-61 sequence and expression was not affected by interruption of the CD-61-specific ORF. Expression of the reporter genes was under the control of a transcription-associated sequence (TAS) derived from the Beaudette gene 5, normally used for the transcription of IBV subgenomic mRNA 5. The Beaudette gene 5 TAS is composed of two tandem repeats of the IBV canonical consensus sequence involved in the acquisition of a leader sequence during the discontinuous transcription of IBV subgenomic mRNAs. It is demonstrated that only one canonical sequence is required for expression of mRNA 5 or for the expression of an mRNA from a D-RNA and that either sequence can function as an acceptor site for acquisition of the leader sequence.
-
-
-
Gill-associated virus of Penaeus monodon prawns: an invertebrate virus with ORF1a and ORF1b genes related to arteri- and coronaviruses
More LessA 20089 nucleotide (nt) sequence was determined for the 5′ end of the (+)-ssRNA genome of gill-associated virus (GAV), a yellow head-like virus infecting Penaeus monodon prawns. Clones were generated from a ∼22 kb dsRNA purified from lymphoid organ total RNA of GAV-infected prawns. The region contains a single gene comprising two long overlapping open reading frames, ORF1a and ORF1b, of 4060 and 2646 amino acids, respectively. The ORFs are structurally related to the ORF1a and ORF1ab polyproteins of coronaviruses and arteriviruses. The 99 nt overlap between ORF1a and ORF1b contains a putative AAAUUUU ‘slippery’ sequence associated with −1 ribosomal frameshifting. A 131 nt stem–loop with the potential to form a complex pseudoknot resides 3 nt downstream of this sequence. Although different to the G/UUUAAAC frameshift sites and ‘H-type’ pseudoknots of nidoviruses, in vitro transcription/translation analysis demonstrated that the GAV element also facilitates read-through of the ORF1a/1b junction. As in coronaviruses, GAV ORF1a encodes a 3C-like cysteine protease domain located between two hydrophobic regions. However, its sequence suggests some structural relationship to the chymotrypsin-like serine proteases of arteriviruses. ORF1b encodes homologues of the ‘SDD’ polymerase, which among (+)-RNA viruses is unique to nidoviruses, as well as metal-ion-binding and helicase domains. The presence of a dsRNA replicative intermediate and ORF1a and ORF1ab polyproteins translated by a −1 frameshift suggests that GAV represents the first invertebrate member of the Order Nidovirales.
-
-
-
Leader switching occurs during the rescue of defective RNAs by heterologous strains of the coronavirus infectious bronchitis virus
More LessA defective RNA (D-RNA), CD-61, derived from the Beaudette strain of the avian coronavirus infectious bronchitis virus (IBV), was rescued (replicated and packaged) using four heterologous strains of IBV as helper virus. Sequence analysis of the genomic RNA from the four heterologous IBV strains (M41, H120, HV10 and D207) identified nucleotide differences of up to 17% within the leader sequence and up to 4·3% within the whole of the adjacent 5′ untranslated region (UTR). Analysis of the 5′ ends of the rescued D-RNAs showed that the Beaudette leader sequence, present on the initial CD-61, had been replaced with the corresponding leader sequence from the helper IBV strain but the adjacent 5′ UTR sequence of the rescued D-RNAs corresponded to the original CD-61 Beaudette sequence. These results demonstrated that the phenomenon of leader switching previously identified for the coronaviruses murine hepatitis virus and bovine coronavirus (BCoV) also occurred during the replication of IBV D-RNAs. Three predicted stem–loop structures were identified within the 5′ UTR of IBV. Stem–loop I showed a high degree of covariance amongst the IBV strains providing phylogenetic evidence that this structure exists and is potentially involved in replication, supporting previous observations that a BCoV stem–loop homologue was essential for replication of BCoV defective interfering RNAs.
-
-
-
Characterization of the sialic acid binding activity of transmissible gastroenteritis coronavirus by analysis of haemagglutination-deficient mutants
More LessTransmissible gastroenteritis coronavirus (TGEV) agglutinates erythrocytes of several species by virtue of sialic acid binding activity of the surface protein S. We have isolated and characterized five haemagglutination-defective (HAD) mutants. In contrast to the parental virus, the mutants were unable to bind to porcine submandibulary mucin, a substrate rich in sialic acid. Each of the mutants was found to contain a single point mutation in the S protein (Cys155Phe, Met195Val, Arg196Ser, Asp208Asn or Leu209Pro), indicating that these amino acids are affecting the sialic acid binding site. In four of the HAD mutants a nearby antigenic site is affected in addition to the sialic acid binding site, as indicated by reactivity with monoclonal antibodies. The parental virus was found to have an increased resistance to the detergent octylglucoside compared to the HAD mutants. This effect depended on cellular sialoglycoconjugates bound to the virion. If the binding of sialylated macromolecules was prevented by neuraminidase treatment, the parental virus was as sensitive to octylglucoside as were the HAD mutants. We discuss the possibility that the sialic acid binding activity helps TGEV to resist detergent-like substances encountered during the gastrointestinal passage and thus facilitates the infection of the intestinal epithelium. An alternative function of the sialic acid binding activity – accessory binding to intestinal tissues – is also discussed.
-
-
-
Clearance of infection in cats naturally infected with feline coronaviruses is associated with an anti-S glycoprotein antibody response
More LessWe have investigated by Western blotting the antibody responses against the three major structural proteins in cats naturally infected with feline coronaviruses that cleared virus infection (group I), established chronic asymptomatic infection (group II) or were sick (group III). The cats of group I developed an anti-S glycoprotein response that was, relative to the anti-M glycoprotein response, at least 30-fold higher than that of chronically infected cats from groups II and III. These results suggest that the anti-S glycoprotein response against antigenic domains revealed by Western blot is associated with clearance of the virus after natural infection, and is not a risk factor for the establishment of a chronic infection.
-
-
-
Identification and subcellular localization of a 41 kDa, polyprotein 1ab processing product in human coronavirus 229E-infected cells.
More LessThe translation products of the human coronavirus (HCV) 229E open reading frames 1a and 1b, the polyproteins 1a and 1ab, are processed by virus- encoded proteinases. One of the key enzymes in this process is a chymotrypsin-like enzyme, the 3C- like proteinase. In this study we have identified an ORF 1b-encoded, 41 kDa processing product in HCV 229E-infected cells by using a monoclonal antibody with defined specificity. We show that this polypeptide is released from polyprotein 1ab by 3C-like proteinase-mediated cleavage of the peptide bonds Gln-6110/Gly-6111 and Gln-6458/Ser- 6459. Also, we have investigated the subcellular localization of the 41 kDa processing product. Immunofluorescence microscopy revealed a punctate, perinuclear distribution of the 41 kDa polypeptide in infected cells and an identical subcellular localization was observed for three additional pp1ab-derived polypeptides. In contrast, the virus nucleocapsid protein showed a homogeneous cytoplasmic localization.
-
-
-
Identification of residues critical for the human coronavirus 229E receptor function of human aminopeptidase N.
More LessAminopeptidase N (APN) is the major cell surface receptor for group 1 coronaviruses. In this study, we have isolated and characterized a feline APN cDNA and shown that the transfection of human embryonic kidney cells with this cDNA renders them susceptible to infection with the feline coronavirus feline infectious peritonitis virus, the human coronavirus (HCV) 229E and the porcine coronavirus porcine transmissible gastroenteritis virus. By using chimeric APN genes, assembled from porcine and feline sequences, we have shown that, analogously to the human APN protein, a region within the amino-terminal part of the feline APN protein (encompassing amino acids 132–295) is essential for its HCV 229E receptor function. Furthermore, by comparing the relevant feline, human and porcine APN sequences, we were able to identify a hypervariable stretch of eight amino acids that are more closely related in the feline and human APN proteins than in the porcine APN molecule. Using PCR- directed mutagenesis, we converted this stretch of amino acids within the porcine APN molecule to the corresponding residues of the human APN molecule. These changes were sufficient to convert porcine APN into a functional receptor for HCV 229E and thus define a small number of residues that are critically important for the HCV 229E receptor function of human APN.
-
-
-
No evidence for quasispecies populations during persistence of the coronavirus mouse hepatitis virus JHM: sequence conservation within the surface glycoprotein gene S in Lewis rats
More LessThe surface glycoprotein S (spike) of coronaviruses is believed to be an important determinant of virulence and displays extensive genetic polymorphism in cell culture isolates. This led us to consider whether the observed heterogeneity is reflected by a quasispecies distribution of mutated RNA molecules within the infected organ. Corona- virus infection of rodents is a useful model system for investigating the pathogenesis of virus-induced central nervous system (CNS) disease. Here, we investigated whether genetic changes in the S gene occurred during virus persistence in vivo. We analysed the variability of S gene sequences directly from the brain tissue of Lewis rats infected with the coronavirus mouse hepatitis virus (MHV) variant JHM-Pi using RT-PCR amplification methods. The S gene sequence displayed a remarkable genetic stability in vivo. No evidence for a quasispecies distribution was found by sequence analysis of amplified S gene fragments derived from the CNS of Lewis rats. Furthermore, the S gene also remained conserved under the selection pressure of a neutralizing antibody. Only a few mutations predicted to result in amino acid changes were detected in single clones. The changes were not represented in the consensus sequence. These results indicate that to retain functional proteins under the constraints of a persistent infection in vivo, conservation of sequence can be more important than heterogeneity.
-
-
-
Expression and characterization of a recombinant murine coronavirus 3C-like proteinase
More LessThe replication of coronaviruses involves proteolytic processing of the gene 1 translation products, pp1a and pp1ab. One of the key enzymes in this process is predicted to be a virus-encoded 3C-like proteinase. In this report, we describe a bacterial system that has allowed us to express and characterize a recombinant murine coronavirus (MHV- JHM) 3C-like proteinase. The partially purified protein has been shown to exhibit proteolytic activity in trans and mutation analysis has been used to demonstrate the indispensability of Cys- 3495 for enzymatic activity. Finally, the effect of class-specific proteinase inhibitors on the trans cleavage activity of the MHV 3C-like proteinase has been used to demonstrate the functional and structural homology of this enzyme to the picorna- virus 3C proteinases.
-
-
-
Characterization of functional domains in the human coronavirus HCV 229E receptor
More LessHuman aminopeptidase N (hAPN or CD13) and porcine aminopeptidase N (pAPN) are functional receptors for human coronavirus (HCV) 229E and porcine transmissible gastroenteritis virus (TGEV), respectively. However, hAPN cannot function as a receptor for TGEV and pAPN cannot function as a receptor for HCV 229E. In this study, we constructed a series of chimeric hAPN/pAPN genes and expressed the corresponding proteins in transfected cells. Subsequently, we identified the chimeric proteins that can function as a receptor for HCV 229E. The results show that replacement of a small region of pAPN sequence (pAPN amino acids 255–348) with the corresponding hAPN sequence (hAPN amino acids 260–353) converts pAPN into a functional receptor for HCV 229E. The region of hAPN that we have defined in this way does not correspond to the region of pAPN that has been identified as essential for the TGEV-receptor interaction. We conclude that although both viruses use a homologous receptor protein, different regions of the protein are required to mediate susceptibility to infection with HCV 229E and TGEV.
-
-
-
Virus entry into a polarized epithelial cell line (MDCK): similarities and dissimilarities between influenza C virus and bovine coronavirus
More LessWe have analysed the uptake of influenza C virus and bovine coronavirus (BCV) by a polarized epithelial cell line, Madin-Darby canine kidney (MDCK) cells. Both viruses use N-acetyl-9-O-acetyl-neuraminic acid as a receptor determinant for attachment to cells. Virus binding assays with immobilized proteins indicated that a glycoprotein of 40 kDa is the major surface protein containing the receptor determinant for the two viruses. MDCK cells grown on filters for permeable support were found to have differential sensitivity to infection by these viruses. Both viruses were able to initiate infection via the apical domain of the plasma membrane, but only influenza C virus also accomplished infection via the basolateral plasma membrane. The resistance of MDCK cells to BCV infection from the basal filter chamber was overcome when the cell polarity was abolished by maintaining the cells in calcium-free medium. This finding indicates that the resistance to basolateral infection by BCV is a property of the cell line and not due to a technical problem related to the use of filters. Our results indicate that two viruses which use the same receptor for attachment to cells may differ in their ability to enter polarized cells. The possible involvement of an accessory molecule in the entry of BCV is discussed.
-
-
-
The region between the M and S genes of porcine haemagglutinating encephalomyelitis virus is highly similar to human coronavirus OC43
More LessThe nucleotide sequences of the regions between the membrane and spike protein genes of three strains of porcine haemagglutinating encephalomyelitis virus (HEV) were determined. A total of 739 (HEV strain 67N) and 751 (strains NT9 and VW572) nucleotides were sequenced. Two ORFs, potentially encoding proteins of 12.8 and 9.6 kDa, were identified. Pairwise comparisons with the corresponding ORFs in bovine coronavirus (BCV) and human coronavirus (HCV) OC43 revealed sequence similarities of greater than 88.5% at the nucleotide and 85.3% at the amino acid level for the 12.8 kDa ORF product. For the 9.6 kDa ORF product similarities were greater than 96.9% and 95.2%, respectively. An additional 12 nucleotide deletion upstream of the 12.8 kDa ORF start codon was found in HEV 67N compared to NT9 and VW572. These results reveal a genomic organization of HEV in the region analysed that is homologous to HCV OC43 but different from BCV.
-
-
-
An adenovirus recombinant expressing the spike glycoprotein of porcine respiratory coronavirus is immunogenic in swine
More LessThe full-length spike (S) gene of porcine respiratory coronavirus (PRCV) was inserted into the genome of human adenovirus type 5 downstream of the early transcription region 3 promoter. The recombinant virus replicated in cultures of the swine testicle ST cell line and directed the synthesis of S antigen with a maximum yield of approximately 26 µg per 106 cells. The antigen was cell-associated except in the late phase of the infection, when a small amount (3.5 µg per 106 cells) was released. The cell-associated antigen consisted of polypeptides of molecular mass 160 kDa and 175 kDa, comigrating with the authentic precursor S′ and the mature S protein of PRCV, respectively. The extracellular recombinant antigen corresponded to the 175 kDa mature protein. Some recombinant S protein was exposed on the cell surface and was recognized by neutralization-mediating anti-S monoclonal antibodies. Piglets, inoculated oronasally with the recombinant adenovirus vector developed PRCV-neutralizing serum antibodies and were partially protected against PRCV challenge, demonstrating the potential of live adenovirus as vaccine vector.
-
-
-
A region of the coronavirus infectious bronchitis virus 1a polyprotein encoding the 3C-like protease domain is subject to rapid turnover when expressed in rabbit reticulocyte lysate
More LessIn order to investigate the mechanisms involved in the processing of infectious bronchitis virus polyproteins, several candidate regions of the genome have been cloned and expressed in vitro. During these studies it was observed that the translation product encoded by one of these clones (pKT205) was poorly expressed. Biochemical and genetic analyses revealed that the basis for the poor expression was a post-translational event involving ubiquitination of the protein and degradation by an ATP-dependent system operating in the reticulocyte lysate used for the in vitro expression. Two independently acting regions which conferred instability were identified, one of which mapped to the predicted 3C protease domain, contained within the 5′ end of the clone, while the other, more C-terminal region, was effective in conferring instability upon a heterologous protein to which it had been transferred. These regions may influence the stability of the authentic viral protein(s) in vivo and hence allow for the control of their expression and/or function at the level of proteolysis by cellular protease(s).
-
-
-
Comparison of bovine coronavirus isolates associated with neonatal calf diarrhoea and winter dysentery in adult dairy cattle in Québec
More LessCytopathic coronaviruses were isolated in HRT-18 cells from bloody faecal samples collected from cows in Québec dairy herds having experienced typical outbreaks of winter dysentery (WD). The formation of polykaryons in the infected cell cultures was found to be dependent on the presence of trypsin in the medium. The WD isolates differed from the prototype Mebus strain of bovine enteropathogenic coronavirus (BCV.Meb) in respect to haemagglutination inhibition (HI), haemagglutination patterns at 4 °C and 37 °C, and receptor destroying enzyme activity with rat erythrocytes. Other field strains of BCV associated with outbreaks of neonatal calf diarrhoea (NCD) also differed from the BCV.Meb strain by demonstrating differences in HI. In all cases, no differences were detected by virus neutralization and Western immunoblotting. Analysis and comparison of the nucleotide and deduced amino acid sequences of the PCR-amplified haemagglutinin esterase (HE) genes of one representative WD strain (BCQ.2590) and two highly cytopathic NCD strains (BCQ.3 and BCQ.571) revealed high degrees of similarities (nt and aa sequence homologies > 98%) with the BCV.Meb strain. The putative esterase active site FGDS was conserved among these four BCV strains, indicating that this domain is probably not a determinant for BCV virulence. Six amino acid substitutions occurred between the HE glycoproteins of BCV.Meb and BCQ.2590 strains; two proline substitutions occurred respectively in the signal peptide (at aa 5) and near the sequences of the putative esterase domain (at aa 53).
-
-
-
Sequence and expression of the ns2 protein gene of human coronavirus OC43
More LessThe complete nucleotide sequence of the ns2 gene of human coronavirus OC43 (HCV-OC43) was determined. Sequence analysis revealed an open reading frame that could encode a protein of 278 amino acids, with an estimated molecular mass of 32.2 kDa. Six potential phosphorylation sites are present but no sites of N-glycosylation were found. The amino acid sequence of the HCV-OC43 ns2 protein shows 92% identity with that of the Mebus strain of bovine coronavirus (BCV). However, a stretch of nine consecutive amino acids near the C terminus is completely different, causing it to be very hydrophilic, which contrasts with the hydrophobic nature of this region in BCV. As shown by immunofluorescence with a monospecific antiserum, the ns2 protein was expressed in the cytoplasm of HCV-OC43-infected HRT-18 cells.
-
