Coronaviruses
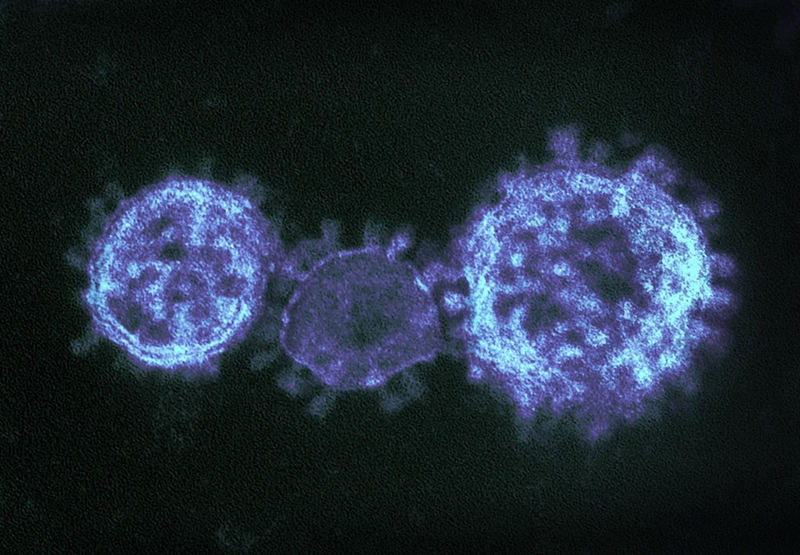
Coronaviruses are a large family of viruses that can infect a range of hosts. They are known to cause diseases including the common cold, Severe Acute Respiratory Syndrome (SARS) and Middle East Respiratory Syndrome (MERS) in humans.
In January 2020, China saw an outbreak of a new coronavirus strain now named SARS-CoV-2. Although the animal reservoir for the SARS and MERS viruses are known, this has yet to have been confirmed for SARS-CoV-2. All three strains are transmissible between humans.
To allow the widest possible distribution of relevant research, the Microbiology Society has brought together articles from across our portfolio and made this content freely available.
Image credit: "MERS-CoV" by NIAID is licensed under CC BY 2.0, this image has been modified.
Collection Contents
21 - 40 of 40 results
-
-
Characterization of the IgA and subclass IgG responses to neutralizing epitopes after infection of pregnant sows with the transmissible gastroenteritis virus or the antigenically related porcine respiratory coronavirus
More LessIn this study, we have investigated the characteristics of secreted IgA and other classes of Ig induced after vaccination of sows with transmissible gastroenteritis virus (TGEV) or the antigenically related porcine respiratory coronavirus (PRCV). Both viruses induced the secretion of neutralizing antibodies of different classes in the sows’ milk, but these protected suckling piglets against TGEV to different degrees. Quantitative differences in the induction of IgA by both viruses were found among the different viral antigenic sites and subsites of glycoprotein S. In TGEV-vaccinated sows, antigenic subsite A was the best inducer of IgA, followed by antigenic site D. After vaccination with PRCV, lower levels of IgA were detected on colostrum and milk, antigenic site D and subsite Ab being the immunodominant sites. This quantitative difference in epitope recognition could explain the differences in newborn piglet protection found using Ig classes purified from the milk of sows immunized with both viruses. Apparently only IgA recognizing at least antigenic sites A and D confers good protection in vivo, whereas any Ig class recognizing only one antigenic site may neutralize the virus in cell culture. These results indicate that the formulation of a subunit vaccine against TGEV has to consider the inclusion of more than one antigenic site involved in virus neutralization.
-
-
-
Cloning and in vitro Expression of the Gene for the E3 Haemagglutinin Glycoprotein of Bovine Coronavirus
More LessSUMMARYA cDNA clone representing the gene for the E3 glycoprotein, the haemagglutinin, of bovine coronavirus was isolated from a plasmid cDNA library of the viral genome and sequenced. The gene is located immediately 5′ of the E2 glycoprotein gene on the viral genome. Nucleotide sequencing of the E3 gene predicts a polypeptide of 424 amino acids with an M r of 47K. In vitro translation of mRNA transcribed from the cloned E3 gene yielded a polypeptide of M r 45K, similar to that predicted from the nucleotide sequence. In the presence of microsomal membranes, the in vitro product was cotranslationally processed to a 62K polypeptide which comigrated on SDS–polyacrylamide gels with the E3 monomer (gp62) obtained from virus-infected cells. Both the 45K and 62K polypeptides were immunoprecipitated with E3-specific monoclonal antibodies, confirming the identity of the gene as that encoding the E3 glycoprotein. Finally, only monoclonal antibodies to the E3 protein inhibited haemagglutination by the virus thus confirming its identity as the haemagglutinin of bovine coronavirus.
-
-
-
Completion of the Sequence of the Genome of the Coronavirus Avian Infectious Bronchitis Virus
More LessSUMMARYThe nucleotide sequence determination of the genome of the Beaudette strain of the coronavirus avian infectious bronchitis virus (IBV) has been completed. The complete sequence has been obtained from 17 overlapping cDNA clones, the 5′-most of which contains the leader sequence (as determined by direct sequencing of the genome) and the 3′-most of which contains the poly(A) tail. Approximately 8 kilobases at the 3′ end of this sequence have already been published. These contain the sequences of mRNAs A to E within which are the genes for the spike, the membrane and the nucleocapsid polypeptides: the main structural components of the virion. The remainder of the sequence, equivalent to the ‘unique’ region of mRNA F, is some 20 kilobases in length and is thought to code for a polymerase or polymerases which are involved in the replication of the genome and the production of the subgenomic messenger RNAs. This sequence contains two large open reading frames, potentially coding for polypeptides of molecular weights 441000 and 300000. Unlike other large open reading frames in the virus, the 300000 open reading frame appears to have no subgenomic RNA associated with it which would allow it to be at the 5′ end of an mRNA species. Because of this, and because of the characteristics of the sequence in the region immediately upstream of its start codon, other mechanisms of translation, such as ribosome slippage, must be postulated.
-
-
-
Comparison of the Spike Precursor Sequences of Coronavirus IBV Strains M41 and 6/82 with that of IBV Beaudette
More LessSummaryThe nucleotide sequences of the spike precursor genes of infectious bronchitis virus strains M41 and 6/82 have been determined and compared with that of the Beaudette strain which we have previously sequenced. The two Massachusetts strains, M41 and Beaudette, were found to be remarkably similar, having only 3.7% of the amino acids different. The situation with 6/82, one of the new field isolates, is quite different and this strain had 13.8% of its amino acids different from Beaudette. The differences identified are discussed in terms of the structural features of the spike protein.
-
-
-
Coronavirus IBV: Removal of Spike Glycopolypeptide S1 by Urea Abolishes Infectivity and Haemagglutination but Not Attachment to Cells
More LessSummaryUrea has been used to remove the S1 spike glycopolypeptide from avian infectious bronchitis virus (IBV) strains M41 and Beaudette, without removing the S2 spike-anchoring glycopolypeptide. Reduction of the pH to 2.9 did not cause release of S1 although some S1 was released spontaneously from IBV Beaudette at pH 7.4. Virus that lacked S1 was no longer infectious or able to cause haemagglutination (HA). However, radiolabelled IBV that lacked S1 attached to erythrocytes and chick kidney cells to the same or similar extent as did intact virus. Treatment of IBV with a phospholipase C preparation, required to make IBV cause HA, did not increase binding of IBV to erythrocytes. The results indicate that while the attachment to cells of virus that lacks S1 is qualitatively different from that of intact virus, the decline in infectivity is the consequence of the loss of some other spike function.
-
-
-
Coronavirus IBV: Virus Retaining Spike Glycopolypeptide S2 but Not S1 Is Unable to Induce Virus-neutralizing or Haemagglutination-inhibiting Antibody, or Induce Chicken Tracheal Protection
More LessSummaryAvian infectious bronchitis coronavirus (IBV) inactivated by β-propiolactone induce partial protection of the trachea in up to 40% of chickens following one intramuscular inoculation 4 to 6 weeks prior to challenge. Retention of an intact tracheal ciliated epithelium 4 days after challenge was the criterion of protection. There was no correlation between protection and serum titres of virus-neutralizing (VN) and haemagglutination-inhibiting (HI) antibody, which were maximal at about 4 weeks after inoculation. Virus from which the S1 but not the S2 (spike-anchoring) spike glycopolypeptide had been removed by urea did not induce protection or VN or HI antibody. Four intramuscular inoculations of monomeric S1 induced VN and HI antibody in two and four chickens respectively. These results indicate that VN and HI antibodies are induced primarily by S1, that intact spikes are a major requirement for the induction of protective immunity and that this propertyis probably associated with S1.
-
-
-
Chronic Shedding of Bovine Enteric Coronavirus Antigen-Antibody Complexes by Clinically Normal Cows
More LessSUMMARYUsing an ELISA for the detection of virus-specific immune complexes, ten cows were found to be shedding bovine enteric coronavirus. The shedding patterns from five of these animals were followed for a period of 12 weeks, and all were found to be chronically shedding virus. Despite the presence of both faecal and serum antibody the infection was not cleared; therefore, the role of cell-mediated immunity (CMI) was investigated by immunosuppressing the chronically shedding cows with dexamethasone. No major role for CMI in maintaining the chronic infection could be determined, although immunosuppression did result in a temporary reduction in the shedding of virus-specific immune complexes.
-
-
-
Cloning and Sequencing of the Gene Encoding the Spike Protein of the Coronavirus IBV
More LessSUMMARYRNA sequences encoding the surface projection (spike) of the coronavirus infectious bronchitis virus, strain Beaudette, have been cloned into pBR322 using cDNA primed with a specific oligonucleotide. A 5.3 kilobase viral insert in the clone pMB179 has been identified. The region of this clone coding for the spike gene has been sequenced by the chain termination method, and we present here the first report of DNA sequence data for a coronavirus spike protein, the protein which forms the characteristic ‘corona’ after which the group is named. The amino acid sequence of the primary translation product, deduced from the DNA sequence, predicts a polypeptide of 1162 amino acids with a molecular weight of 127006. This has many interesting features which confirm and extend our knowledge of this recently characterized membrane glycoprotein. The polypeptide is subsequently cleaved to S1 and S2, and partial amino acid analysis of the amino-terminus of the S1 polypeptide has been employed to locate the position of this terminus of S1 within the large open reading frame. The amino acid analysis also reveals the presence of an 18 amino acid putative signal sequence on the primary translation product which is not present on the mature S1 polypeptide.
-
-
-
Coding Sequence of Coronavirus MHV-JHM mRNA 4
More LessSUMMARYA coding sequence at the 5′ end of mRNA 4 of the coronavirus MHV-JHM was determined by M13/chain-terminator sequencing of cloned cDNA. An open reading frame of 417 bases with the potential to encode a polypeptide of mol. wt. 15200 (139 residues) was identified. The 3′ end of the open reading frame overlapped by 16 bases the start of an open reading frame found in mRNA 5. The translation product of mRNA 4 was predicted to be a basic polypeptide rich in threonine. It had a large hydrophobic region near the amino terminus and a basic carboxy terminus. An intracellular, virus-specific polypeptide, which has been previously described as having a mol. wt. of 14000 to 14500 has the size and charge characteristics of such a translation product.
-
-
-
Coronavirus MHV-JHM mRNA 5 Has a Sequence Arrangement which Potentially Allows Translation of a Second, Downstream Open Reading Frame
More LessSUMMARYThe sequence of a 5′-proximal region of mRNA 5 of coronavirus MHV-JHM was determined by chain-terminator sequencing of cDNA subcloned in M13. The sequence contained two long open reading frames of 321 bases and 264 bases, overlapping by five bases but in different frames. Both open reading frames are initiated by AUG codons in sequence contexts that are relatively infrequently used as initiator codons. The smaller, downstream open reading frame encoded a neutral protein (mol. wt. 10200) with a hydrophobic amino terminus. The larger, 5′-proximal open reading frame encoded a basic protein (mol. wt. 12400) which lacks internal methionine residues. With the exception of the AUG codon initiating the downstream open reading frame, no internal AUG codons were found within the sequence covered by the upstream open reading frame. These results suggest that the MHV-JHM mRNA 5 is translated to produce two proteins by a mechanism involving internal initiation of protein synthesis. Preliminary evidence is presented showing that the downstream open reading frame is functional in vivo.
-
-
-
Coronavirus IBV: Structural Characterization of the Spike Protein
More LessSUMMARYThe spike protein (S; surface projection) of avian infectious bronchitis virus (IBV) strain M41 comprises two glycopolypeptides, S1 (mol. wt. 90 × 103) and S2 (mol. wt. 84 × 103), in equimolar proportions. The apparent mol. wt. of S was calculated as 354 (±17) × 103 following co-sedimentation with catalase in sucrose gradients. Incubation of radiolabelled IBV with urea resulted in the removal of most S1, but none of S2, from the virus particle. A similar result was obtained using low concentrations of SDS, although some nucleocapsid, but not matrix, protein was also released. 2% SDS alone was as effective as 2% SDS plus 2% 2-mercaptoethanol for the separation of S1 and S2 prior to SDS–polyacrylamide gel electrophoresis. Dithiothreitol did not remove S from virions but did decrease the buoyant density of the virus from 1.18 g/ml to 1.16 g/ml, and changed the configuration of S. It is concluded that IBV S protein is an oligomer comprising two copies of each of S1 and S2, although the possibility that there are three copies of each glycopolypeptide cannot be discounted. S is attached to the membrane by S2, while S1 has little or no contact with the membrane and may form the major part of the bulbous end of S. Interpeptide disulphide bonds do not occur in S, and the association of S1 and S2 is weak.
-
-
-
Coronavirus IBV: Further Evidence that the Surface Projections are Associated with Two Glycopolypeptides
More LessSUMMARYThe surface projections (peplomers) of avian infectious bronchitis virus (IBV) strain M41 have been separated from the nucleocapsid (N) and matrix (M) proteins by sedimentation in a sucrose gradient after virus disruption by the non-ionic detergent Nonidet P40. The peplomers comprised two glycopolypeptides of mol. wt. 90 × 103 (90K; S1) and 84K (S2), shown by analysis of differentially radiolabelled virus to be present in equimolar proportions. Polypeptides of 75K and 110K, which were detected by Coomassie Brilliant Blue staining in similar amounts to S1 and S2 in some unlabelled virus preparations, were absent from peplomer preparations and are probably host cell polypeptides. The S1:S2:N:M polypeptide molar ratio for IBV-M41 was approximately 1:1:6:15.
-
-
-
Coronavirus IBV Glycopolypeptides: Size of Their Polypeptide Moieties and Nature of Their Oligosaccharides
More LessSUMMARYAnalysis of differentially radiolabelled avian infectious bronchitis virus (IBV) indicated that the matrix (M) polypeptides of mol. wt. 23 × 103 (23K), 26K, 28K, 30K and 34K (M23 to M34) which have been shown to give the same peptide maps, differed in their degree of glycosylation; M23 was not glycosylated while glycosylation increased with increasing mol. wt. from M26 to M34. Both glucosamine and mannose were components of M26 to M34 but [3H]fucose appeared to be associated mainly with M34. Endo-β-N-acetylglucosaminidase H removed oligosaccharides from M28 and M30 but not M26 and M34, to give a polypeptide of 23K. The surface projection glycopolypeptides S1 (90K) and S2 (84K) incorporated 3H-labelled glucosamine and mannose but not fucose and had oligosaccharides removed by endoglycosidase H. The mol. wt. of the resultant polypeptides varied among experiments; the lowest mol. wt. observed were 64K and 61K. These results indicate (i) that the polypeptide moieties of the S polypeptides are approximately 64K and 61K, and 23K for the M polypeptide, (ii) that the oligosaccharides of the S and M polypeptides are of the high-mannose type and are linked to the polypeptides by N-glycosidic linkages, and (iii) that the M glycoprotein of IBV differs from that of murine coronaviruses and bovine coronavirus L9 which have O-linked oligosaccharides.
-
-
-
Characterization of Murine Coronavirus RNA by Hybridization with Virus-specific cDNA Probes
More LessSUMMARYGenome RNA of mouse hepatitis virus (MHV) strain A59 has been used as a template to synthesize two virus-specific probes: cDNArep, representing the majority of sequences of the genome RNA and cDNA3′, representing the 3′ end of the genome RNA. Molecular hybridization with these cDNAs was used to characterize both genome RNA and intracellular virus-specific RNAs. Hybridization of genome RNAs of MHV strains A59, JHM, and MHV-3 with A59 cDNArep showed that, although these three strains exhibit different pathogenicities, they contain closely related nucleotide sequences. Hybridization of intracellular RNA from MHV-infected cells with virus-specific cDNA shows that (i) the majority of virus-specific RNA is polyadenylated, (ii) virus-specific intracellular RNA contains six subgenomic species of the same polarity as genome RNA and (iii) all subgenomic RNAs contain the same 3′ sequences as the genome RNA and thus form a nested set of RNAs.
-
-
-
Coronavirus JHM: Coding Assignments of Subgenomic mRNAs
More LessSUMMARYProtein synthesis in the murine hepatitis virus JHM-infected cells was temporarily inhibited by hypertonic shock. When the cells were returned to isotonic medium the synthesis of six virus-specific polypeptides, 150K, 65K, 60K, 30K, 23K and 14K was reinitiated simultaneously. Polyadenylated RNA isolated from the cytoplasm or polysomes of infected cells was translated in vitro and the products included polypeptides with molecular weights (mol. wt.) of 120000, 60000, 30000, 23000 and 14000. Immunoprecipitation and fingerprinting of [35S]methionine-containing tryptic peptides showed that the 60000 and 23000 mol. wt. products were identical to the 60K and 23K polypeptides found in infected cells; the 120000 mol. wt. product showed identity with the 150K intracellular polypeptide and a virus-specific 120K polypeptide synthesized in tunicamycin-treated cells. Two-dimensional polyacrylamide gel electrophoresis strongly suggested that the 30000 and 14000 mol. wt. products are equivalent to virus-specific 30K and 14K intracellular polypeptides. [3H]Uridine-labelled polyadenylated virus RNA was isolated from infected cells and sedimented in sucrose gradients containing formamide. The distribution in the gradient of each of the previously identified virus RNAs was determined by gel electrophoresis and gradient fractions enriched for each RNA were translated in vitro. The 120000, 60000, 30000, 23000 and 14000 mol. wt. polypeptides were found to be encoded by mRNAs 3, 7, 2, 6, and 4 or 5 respectively. These results demonstrate that the virus-specific polypeptides in JHM-infected cells are encoded in separate subgenomic mRNAs and are translated independently. The assignment of coding functions and the known sequence relationships of JHM RNAs permitted a gene order to be deduced.
-
-
-
Coronavirus JHM: Tryptic Peptide Fingerprinting of Virion Proteins and Intracellular Polypeptides
More LessSUMMARYThe virion proteins and intracellular polypeptides of the murine coronavirus MHV-JHM have been analysed by two-dimensional fingerprinting of their [35S]methionine-containing tryptic peptides. The analysis shows that the virion proteins gp98, gp65, pp60 and p23 are distinct. Virion protein gp25 has the same polypeptide component as p23, and virion protein gp170 has a polypeptide component related to gp98. The six virus polypeptides synthesized in infected cells, 150K, 65K, 60K, 30K, 23K and 14K are also distinct. The 170K and 98K species, which are produced by processing, are related to 150K. The 25K species is a processed form of 23K. The identity of corresponding species in the cell and in the virion has been shown and a model describing the genesis of coronavirus JHM proteins can now be proposed.
-
-
-
Coronavirus JHM: Characterization of Intracellular Viral RNA
More LessSUMMARYAfter infection of Sac(-) cells with the murine coronavirus JHM the synthesis of seven major and two minor RNA species was induced. These RNAs were polyadenylated and single-stranded. Their mol. wt. were estimated by electrophoresis in agarose gels containing methylmercury hydroxide. The values for the major species were 6.67 × 106 for RNA of genome size (RNA 1), 3.42 × 106 for RNA 2, 2.76 × 106 for RNA 3, 1.35 × 106 for RNA 4, 1.19 × 106 for RNA 5, 0.93 × 106 for RNA 6 and 0.62 × 106 for RNA 7. The minor species have a size of 4.7 × 106 (RNA a) and 1.5 × 106 (RNA b). The same number of species were found by electrophoresis after denaturation with glyoxal-dimethyl sulphoxide. No gross difference in number of RNAs and the amount of each species was found between total cytoplasmic RNA, polyadenylated cytoplasmic RNA and RNA extracted from pelleted polysomes.
-
-
-
Coronavirus JHM: Intracellular Protein Synthesis
More LessSummaryCoronavirus JHM contained six major proteins, four of which were glycosylated. The proteins were gp170, gp98, gp65, p60, gp25 and p23. Sac(-) cells infected with JHM shut off host cell protein synthesis, and the synthesis of three major (150K, 60K and 23K) and three minor (65K, 30K and 14K) polypeptides was detected by pulse-labelling with 35S-methionine. Antiserum directed against purified virus proteins specifically immunoprecipitated the three major intracellular species and also the 65K minor species. During a chase period, species 150K and 23K were processed and three new immunoprecipitable species, 170K, 98K and 25K appeared. The intracellular species 170K, 98K, 65K, 60K, 25K and 23K co-electrophoresed with virion proteins.
Two-dimensional gel electrophoresis of infected cell polypeptides showed that the 60K, 23K, 25K and 14K species were relatively basic polypeptides whilst the 98K and 170K were relatively acidic and heterogeneously charged polypeptides. Additionally, a charge-size modification of the 23K species during processing was detected, which was not apparent using one-dimensional gel analysis.
-
-
-
Coronavirus JHM: a Virion-associated Protein Kinase
More LessSUMMARYCoronavirus JHM contains six major proteins, one of which, the 60000 mol. wt. nucleocapsid protein pp60, is phosphorylated. In JHM-infected cells ip 60K, the intracellular precursor to pp60 is also phosphorylated. Associated with purified JHM virions is a protein kinase which will phosphorylate pp60 and a variety of exogenous substrates in vitro. The enzyme has the characteristics of a cyclic nucleotide-independent protein kinase. Both the in vivo reaction and the enzyme activity in vitro transferred the γ-phosphate of ATP to serine residues on the nucleocapsid protein.
-
